
John Wiley & Sons - 2004 - Analysis of Genes and Genomes
.pdf

200 CLONING A GENE 5
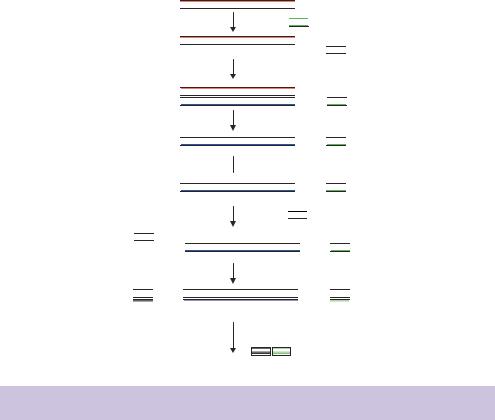
sufficient prior to the progression of the PCR stage, where second-strand cDNA synthesis and subsequent PCR amplification is performed using a thermostable DNA polymerase – e.g. Taq DNA polymerase (see Chapter 4). In addition to their DNA-dependent DNA polymerase activity, some thermostable DNA polymerases (e.g. Thermus thermophilus (Tth) DNA polymerase) possess a reverse transcriptase activity in the presence of manganese ions. This has led to the development of protocols for single-enzyme reverse transcription and PCR amplification (Myers and Gelfand, 1991). Systems have also been developed in which the reverse transcriptase reaction and PCR are performed in the same buffer to eliminate secondary additions to the reaction mix to decrease both hands-on time and the likelihood of introducing contaminants into the reaction (Wang, Cao and Johnson, 1992). Such systems are ideal for the amplification of mRNA molecules whose sequence is already known using highly specific primers, but the construction of an amplified representative library requires additional steps to ensure that each mRNA molecule within the population is represented within the library. Several methods have been devised to amplify all potential mRNA species within a sample. The method outlined in Figure 5.7 utilizes many of the same elements as we have already seen in cDNA library construction. The first cDNA strand is synthesized using reverse transcriptase from an oligo-dT primer to which additional, unique sequences have been added at the 5 -end. The mRNA strand of the RNA –DNA hybrid is removed by treatment with RNaseH, prior to the addition of multiple C residues at the 3 - end of the DNA molecule using terminal transferase. The second cDNA strand is synthesized using an oligo-dG primer that, again, has unique sequences at its 5 -end. The thermostable DNA polymerase that will be used for the subsequent PCR reaction may also be used to perform the synthesis of the second cDNA strand. The unique sequences at the 5 - and 3 -ends of the resulting doublestranded cDNA are then used as primer binding sites for a PCR reaction using primers containing these sequences. The resulting PCR products will contain a huge number of copies of each cDNA molecule produced in the RT reaction.
5.5Subtraction Libraries
As we have discussed earlier, many of the mRNA molecules produced by different cells will be the same. For example, almost all cells need to produce the enzymes required for glucose metabolism, and many of the intracellular protein components of all cells, are identical. Therefore, we might want to just concentrate on the differences between cell types to identify genes that are distinctive to a cell type, developmental stage or particular environmental stress. The advantage of PCR-based cDNA libraries is that they are amenable to
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
5.5 SUBTRACTION LIBRARIES |
201 |
||||||||||||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
Driver cDNA library |
|
|
Tester cDNA library |
|
|||||||||||||||||||||||||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
Boil and anneal
Extract biotin using avidin
|
Amplify using primers |
Selection |
Subtraction |
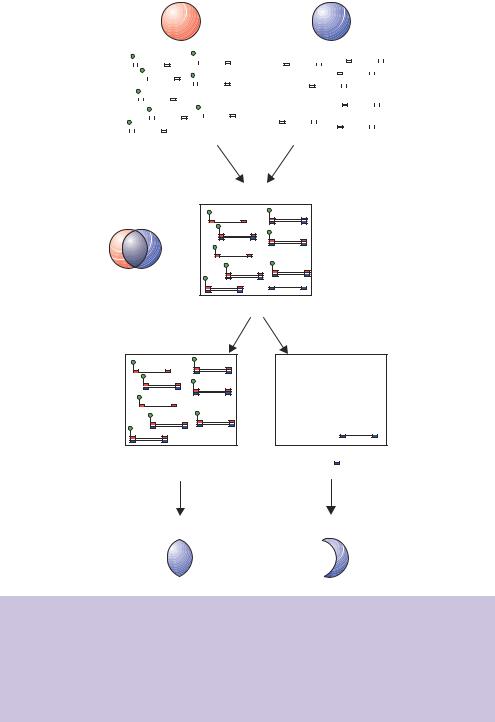
Figure 5.8. Subtractive hybridization of PCR-based libraries. Two PCR-based cDNA libraries are constructed. The driver library is labelled with biotin, while the tester library is not. Hybridization of the two libraries ensures that common sequences will be biotin labelled, while sequences that are unique to the tester library will not. The biotin labelled sequences can be removed from the mixture via binding to avidin, thereby enriching the sequences unique to the tester library. Enriched sequences can be amplified, by PCR, using primers unique to the tester library
5.5 SUBTRACTION LIBRARIES |
203 |
|
|
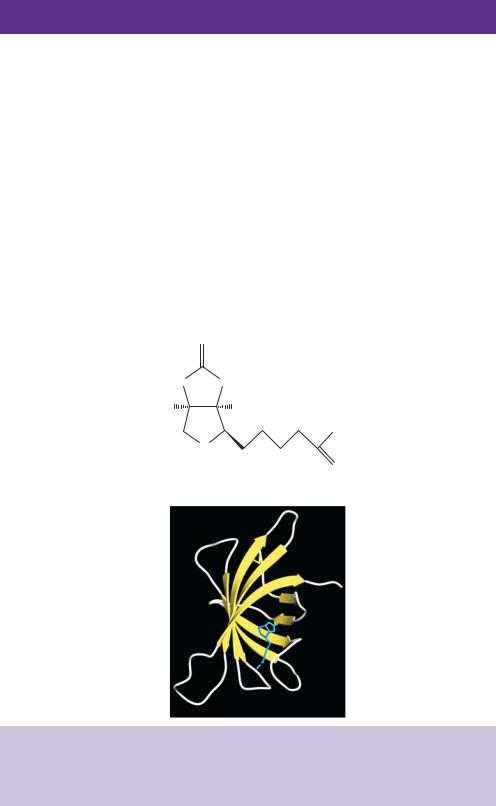
biotin from all but the most dilute of solutions (Green, 1963). Biotin can be chemically coupled to phosphoramidites (see Figure 4.6) through its carboxyl group and thus can be incorporated into chemically synthesized oligonucleotide primers at either the 5 - or 3 -ends or throughout the primer in association with one of the bases. Additionally, biotin may be added to DNA fragments after synthesis using photoactivateable biotin analogues (Barr and Emanuel, 1990). Oligonucleotides and DNA fragments produced that are labelled with biotin have a high affinity for avidin.
The driver cDNA library is denatured, by heating, to separate its individual strands, and then hybridized in large excess to the denatured tester cDNA library (Luqmani and Lymboura, 1994). The majority of the double-stranded cDNA molecules produced by this will be reformed copies of the driver cDNA library since it is present in excess. However, several hybridization possibilities can occur between the driver cDNA and the tester cDNA:
•cDNA molecules only present in the driver cDNA will be unable to bind to a complementary partner from the tester cDNA and will reform as doublestranded DNA with each strand being derived from the driver library.
•cDNA molecules present in both libraries may hybridize to each other to produce hybrids in which one strand is derived from the driver cDNA and one strand is derived from the tester cDNA.
•cDNA molecules only present in the tester cDNA will be unable to bind to a complementary partner from the driver cDNA and will reform as doublestranded DNA, with each strand being derived from the tester library. Importantly, because these molecules contain no driver cDNA, they will not be labelled with biotin. All other DNA species described above will contain at least one DNA strand that possesses a biotin molecule.
Physical separation of the hybrid cDNA molecules by passing the cDNA molecules through a column to which avidin is attached will result in the adherence of all DNA species that possess one or more strands of the driver cDNA. Therefore, only DNA species unique to the tester cDNA library will not adhere to the avidin column. PCR amplification of the column flow-through and the material retained by the column (after elution with free biotin) will result in the formation of a subtraction and a selection library, respectively. The subtraction library contains sequences unique to the tester library that are not present in the driver library, and the selection library contains shared or common sequences that are present in both libraries (Figure 5.10).
Library subtraction is an extremely powerful technique for the enrichment of sequences present only in the tester cDNA library. For example, Brady et al.



206 GENE IDENTIFICATION 6
Here, we will look at each of these methods in turn. Advances in DNA sequencing technology (see Chapter 9) have led to a decrease in the use of nucleic acid hybridization techniques to identify genes, but many of the techniques we will talk about here to identify proteins and protein function have become increasingly popular.
6.1Screening by Nucleic Acid Hybridization
The complementary base pairing of one nucleic acid strand with another can be used to identify recombinant clones that contain DNA sequences that are identical, or similar, to that of a probe sequence. A difficulty that readers will recognize with this type of screening is that you need to know something about the DNA sequence you want to find before a probe can be designed to search for that sequence. We have encountered this problem before when looking at the design of PCR primers (see Chapter 4). Nevertheless, hybridization screening provided the backbone of gene identification for many years. The main advantages of this type of screening are that it does not rely on the expression of the cloned DNA fragments within the library and it can be applied to almost any vector system into which a library has been cloned.
A generalized scheme for the identification of recombinant clones based solely on their DNA sequence is outlined in Figure 6.1. This scheme is based
Nitrocellulose |
|
|
|
Peel sheet to |
Lyse bacteria |
DNA bound |
|
produce replica |
and denature |
||
|
|||
of colonies |
DNA |
|
|
Petri dish with colonies |
|
Incubate |
|
|
with probe |
||
of bacteria containing |
|
||
|
and wash |
||
recombinant plasmids |
|
||
|
|
||
|
Expose |
|
|
|
to film |
|
Figure 6.1. Screening for DNA sequences by nucleic acid hybridization. A sheet of nitrocellulose (or nylon) membrane is placed on top of an agar plate to generate a replica of the bacterial colonies. DNA from the bacteria, which includes the recombinant library, is attached to the membrane after the bacteria have been lysed. The denatured (single-stranded) DNA can then be used as a template for the binding of complementary, radiolabelled, DNA sequences. The binding of these sequences to the membrane can be analysed by exposure of the washed membrane to X-ray film
6.1 SCREENING BY NUCLEIC ACID HYBRIDIZATION |
207 |
|
|
on that originally described by Grunstein and Hogness, but includes several later modifications, which allow the screening of a large number of bacterial colonies from a single plate, and also utilizes a filter lift procedure (Grunstein and Hogness, 1975; Grunstein and Wallis, 1979; Hanahan and Meselson, 1980). The basic idea of this type of screening procedure is to capture the DNA contained within each clone on to a nitrocellulose (or nowadays nylon) filter so that it can be used as a fixed point for the binding of other, complementary, DNA molecules. The colonies to be screened are grown on agar plates that contain the appropriate antibiotics etc. to allow the growth of cells containing the recombinant molecules. When the bacterial colonies have grown, a sheet of nylon is placed on top of them and then lifted off to produce a replica version of the plate. A portion of each bacterial colony will adhere to the filter sheet and will be removed from the agar plate along with the nitrocellulose. The nylon replica is then treated in various ways to lyse the bacteria and firmly attach the DNA to the sheet. Typically, the following steps are carried out:
•the nylon sheet is treated with alkali (e.g. 0.5 M NaOH) to initiate both bacterial cell lysis and DNA denaturation;
•upon neutralization, the sheet is treated with proteases (e.g. proteinase K) in order to remove the protein and leave the denatured DNA bound to the membrane and
•the sheet is then baked at 80 ◦C, or treated with UV light, to firmly adhere the DNA to the membrane.
This results in a nitrocellulose sheet containing a denatured (single-stranded) DNA copy of the bacterial colonies originally present on the agar plate. The single-stranded DNA replica can then be used as a template to bind other DNA molecules. The hybridization of a labelled single-stranded probe to the DNA on the nitrocellulose membrane will reveal the location of colonies on the original dish that contain identical, or at least similar, DNA sequences. The probe can be any single-stranded nucleic acid sequence and does not need to match the target sequence precisely.
An example of a recombinant clone isolated using this approach is the cDNA encoding a protein involved in plasmin production. Plasmin is a potent serine proteinase that has important functions in diverse physiological processes in mammals, such as degradation of extracellular matrix proteins, blood clot dissolution, cellular migration and cancer metastasis (Vassalli and Saurat, 1996). In mammalian plasma, plasmin degrades blood clot networks to produce soluble products. Plasmin is produced from a precursor, plasminogen, through limited proteolysis by plasminogen activators. Plasminogen activators are used


 AAAAA–3'
AAAAA–3'
 TTTTT–3'
TTTTT–3'
 AAAAA–3' 3'–TTTTT
AAAAA–3' 3'–TTTTT
 –5'
–5'
 AAAAA–3' 3'–
AAAAA–3' 3'–
 TTTTT
TTTTT
 –5'
–5'
 TTTTT
TTTTT
 –5'
–5'

 TTTTT
TTTTT
 –5'
–5' 
 GGGGG–3'
GGGGG–3'
 GGGGG–3'
GGGGG–3'
 TTTTT
TTTTT
 –5'
–5' 
 GGGGG
GGGGG
 AAAAA
AAAAA
 –3'
–3' CCCCC
CCCCC
 TTTTT
TTTTT –5'
–5'








