
- •Contents
- •Contributors
- •Part I General Principles of Cell Death
- •1 Human Caspases – Apoptosis and Inflammation Signaling Proteases
- •1.1. Apoptosis and limited proteolysis
- •1.2. Caspase evolution
- •2. ACTIVATION MECHANISMS
- •2.2. The activation platforms
- •2.4. Proteolytic maturation
- •3. CASPASE SUBSTRATES
- •4. REGULATION BY NATURAL INHIBITORS
- •REFERENCES
- •2 Inhibitor of Apoptosis Proteins
- •2. CELLULAR FUNCTIONS AND PHENOTYPES OF IAP
- •3. IN VIVO FUNCTIONS OF IAP FAMILY PROTEINS
- •4. SUBCELLULAR LOCATIONS OF IAP
- •8. IAP–IAP INTERACTIONS
- •10. ENDOGENOUS ANTAGONISTS OF IAP
- •11. IAPs AND DISEASE
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •2.1. The CD95 (Fas/APO-1) system
- •2.1.1. CD95 and CD95L: discovery of the first direct apoptosis-inducing receptor-ligand system
- •2.1.2. Biochemistry of CD95 apoptosis signaling
- •2.2. The TRAIL (Apo2L) system
- •3.1. The TNF system
- •3.1.1. Biochemistry of TNF signal transduction
- •3.1.2. TNF and TNF blockers in the clinic
- •3.2. The DR3 system
- •4. THE DR6 SYSTEM
- •6. CONCLUDING REMARKS AND OUTLOOK
- •SUGGESTED READINGS
- •4 Mitochondria and Cell Death
- •1. INTRODUCTION
- •2. MITOCHONDRIAL PHYSIOLOGY
- •3. THE MITOCHONDRIAL PATHWAY OF APOPTOSIS
- •9. CONCLUSIONS
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •3. INHIBITING APOPTOSIS
- •4. INHIBITING THE INHIBITORS
- •6. THE BCL-2 FAMILY AND CANCER
- •SUGGESTED READINGS
- •6 Endoplasmic Reticulum Stress Response in Cell Death and Cell Survival
- •1. INTRODUCTION
- •2. THE ESR IN YEAST
- •3. THE ESR IN MAMMALS
- •4. THE ESR AND CELL DEATH
- •5. THE ESR IN DEVELOPMENT AND TISSUE HOMEOSTASIS
- •6. THE ESR IN HUMAN DISEASE
- •7. CONCLUSION
- •7 Autophagy – The Liaison between the Lysosomal System and Cell Death
- •1. INTRODUCTION
- •2. AUTOPHAGY
- •2.2. Physiologic functions of autophagy
- •2.3. Autophagy and human pathology
- •3. AUTOPHAGY AND CELL DEATH
- •3.1. Autophagy as anti–cell death mechanism
- •3.2. Autophagy as a cell death mechanism
- •3.3. Molecular players of the autophagy–cell death cross-talk
- •4. AUTOPHAGY, CELLULAR DEATH, AND CANCER
- •5. CONCLUDING REMARKS AND PENDING QUESTIONS
- •SUGGESTED READINGS
- •8 Cell Death in Response to Genotoxic Stress and DNA Damage
- •1. TYPES OF DNA DAMAGE AND REPAIR SYSTEMS
- •2. DNA DAMAGE RESPONSE
- •2.2. Transducers
- •2.3. Effectors
- •4. CHROMATIN MODIFICATIONS
- •5. CELL CYCLE CHECKPOINT REGULATION
- •6. WHEN REPAIR FAILS: SENESCENCE VERSUS APOPTOSIS
- •6.1. DNA damage response and the induction of apoptosis
- •6.2. p53-independent mechanisms of apoptosis
- •6.3. DNA damage response and senescence induction
- •7. DNA DAMAGE FROM OXIDATIVE STRESS
- •SUGGESTED READINGS
- •9 Ceramide and Lipid Mediators in Apoptosis
- •1. INTRODUCTION
- •3.1. Basic cell signaling often involves small molecules
- •3.2. Sphingolipids are cell-signaling molecules
- •3.2.1. Ceramide induces apoptosis
- •3.2.2. Ceramide accumulates during programmed cell death
- •3.2.3. Inhibition of ceramide production alters cell death signaling
- •4.1. Ceramide is generated through SM hydrolysis
- •4.3. aSMase can be activated independently of extracellular receptors to regulate apoptosis
- •4.4. Controversial aspects of the role of aSMase in apoptosis
- •4.5. De novo ceramide synthesis regulates programmed cell death
- •4.6. p53 and Bcl-2–like proteins are connected to de novo ceramide synthesis
- •4.7. The role and regulation of de novo synthesis in ceramide-mediated cell death is poorly understood
- •5. CONCLUDING REMARKS AND FUTURE DIRECTIONS
- •5.1. Who? (Which enzyme?)
- •5.2. What? (Which ceramide?)
- •5.3. Where? (Which compartment?)
- •5.4. When? (At what steps?)
- •5.5. How? (Through what mechanisms?)
- •5.6. What purpose?
- •6. SUMMARY
- •SUGGESTED READINGS
- •1. General Introduction
- •1.1. Cytotoxic lymphocytes and apoptosis
- •2. CYTOTOXIC GRANULES AND GRANULE EXOCYTOSIS
- •2.1. Synthesis and loading of the cytotoxic granule proteins into the secretory granules
- •2.2. The immunological synapse
- •2.3. Secretion of granule proteins
- •2.4. Uptake of proapoptotic proteins into the target cell
- •2.5. Activation of death pathways by granzymes
- •3. GRANULE-BOUND CYTOTOXIC PROTEINS
- •3.1. Perforin
- •3.2. Granulysin
- •3.3. Granzymes
- •3.3.1. GrB-mediated apoptosis
- •3.3.2. GrA-mediated cell death
- •3.3.3. Orphan granzyme-mediated cell death
- •5. CONCLUSIONS
- •REFERENCES
- •Part II Cell Death in Tissues and Organs
- •1.1. Death by trophic factor deprivation
- •1.2. Key molecules regulating neuronal apoptosis during development
- •1.2.1. Roles of caspases and Apaf-1 in neuronal cell death
- •1.2.2. Role of Bcl-2 family members in neuronal cell death
- •1.3. Signal transduction from neurotrophins and neurotrophin receptors
- •1.3.1. Signals for survival
- •1.3.2. Signals for death
- •2.1. Apoptosis in neurodegenerative diseases
- •2.1.4. Amyotrophic lateral sclerosis
- •2.2. Necrotic cell death in neurodegenerative diseases
- •2.2.1. Calpains
- •2.2.2. Cathepsins
- •3. CONCLUSIONS
- •ACKNOWLEDGMENT
- •SUGGESTED READINGS
- •ACKNOWLEDGMENT
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •5. S-NITROSYLATION OF PARKIN
- •7. POTENTIAL TREATMENT OF EXCESSIVE NMDA-INDUCED Ca2+ INFLUX AND FREE RADICAL GENERATION
- •8. FUTURE THERAPEUTICS: NITROMEMANTINES
- •9. CONCLUSIONS
- •Acknowledgments
- •SUGGESTED READINGS
- •3. MITOCHONDRIAL PERMEABILITY TRANSITION ACTIVATED BY Ca2+ AND OXIDATIVE STRESS
- •4.1. Mitochondrial apoptotic pathways
- •4.2. Bcl-2 family proteins
- •4.3. Caspase-dependent apoptosis
- •4.4. Caspase-independent apoptosis
- •4.5. Calpains in ischemic neural cell death
- •5. SUMMARY
- •ACKNOWLEDGMENTS
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •2. HISTORICAL ANTECEDENTS
- •7.1. Activation of p21 waf1/cip1: Targeting extrinsic and intrinsic pathways to death
- •8. CONCLUSION
- •ACKNOWLEDGMENTS
- •REFERENCES
- •16 Apoptosis and Homeostasis in the Eye
- •1.1. Lens
- •1.2. Retina
- •2. ROLE OF APOPTOSIS IN DISEASES OF THE EYE
- •2.1. Glaucoma
- •2.2. Age-related macular degeneration
- •4. APOPTOSIS AND OCULAR IMMUNE PRIVILEGE
- •5. CONCLUSIONS
- •SUGGESTED READINGS
- •17 Cell Death in the Inner Ear
- •3. THE COCHLEA IS THE HEARING ORGAN
- •3.1. Ototoxic hair cell death
- •3.2. Aminoglycoside-induced hair cell death
- •3.3. Cisplatin-induced hair cell death
- •3.4. Therapeutic strategies to prevent hair cell death
- •3.5. Challenges to studies of hair cell death
- •4. SPIRAL GANGLION NEURON DEATH
- •4.1. Neurotrophic support from sensory hair cells and supporting cells
- •4.2. Afferent activity from hair cells
- •4.3. Molecular manifestations of spiral ganglion neuron death
- •4.4. Therapeutic interventions to prevent SGN death
- •ACKNOWLEDGMENTS
- •SUGGESTED READINGS
- •18 Cell Death in the Olfactory System
- •1. Introduction
- •2. Anatomical Aspects
- •3. Life and Death in the Olfactory System
- •3.1. Olfactory epithelium
- •3.2. Olfactory bulb
- •REFERENCES
- •1. Introduction
- •3.1. Beta cell death in the development of T1D
- •3.2. Mechanisms of beta cell death in type 1 diabetes
- •3.2.1. Apoptosis signaling pathways downstream of death receptors and inflammatory cytokines
- •3.2.2. Oxidative stress
- •3.3. Mechanisms of beta cell death in type 2 diabetes
- •3.3.1. Glucolipitoxicity
- •3.3.2. Endoplasmic reticulum stress
- •5. SUMMARY
- •Acknowledgments
- •REFERENCES
- •20 Apoptosis in the Physiology and Diseases of the Respiratory Tract
- •1. APOPTOSIS IN LUNG DEVELOPMENT
- •2. APOPTOSIS IN LUNG PATHOPHYSIOLOGY
- •2.1. Apoptosis in pulmonary inflammation
- •2.2. Apoptosis in acute lung injury
- •2.3. Apoptosis in chronic obstructive pulmonary disease
- •2.4. Apoptosis in interstitial lung diseases
- •2.5. Apoptosis in pulmonary arterial hypertension
- •2.6. Apoptosis in lung cancer
- •SUGGESTED READINGS
- •21 Regulation of Cell Death in the Gastrointestinal Tract
- •1. INTRODUCTION
- •2. ESOPHAGUS
- •3. STOMACH
- •4. SMALL AND LARGE INTESTINE
- •5. LIVER
- •6. PANCREAS
- •7. SUMMARY AND CONCLUDING REMARKS
- •SUGGESTED READINGS
- •22 Apoptosis in the Kidney
- •1. NORMAL KIDNEY STRUCTURE AND FUNCTION
- •3. APOPTOSIS IN ADULT KIDNEY DISEASE
- •4. REGULATION OF APOPTOSIS IN KIDNEY CELLS
- •4.1. Survival factors
- •4.2. Lethal factors
- •4.2.1. TNF superfamily cytokines
- •4.2.2. Other cytokines
- •4.2.3. Glucose
- •4.2.4. Drugs and xenobiotics
- •4.2.5. Ischemia-reperfusion and sepsis
- •5. THERAPEUTIC APPROACHES
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •2. APOPTOSIS IN THE NORMAL BREAST
- •2.1. Occurrence and role of apoptosis in the developing breast
- •2.2.2. Death ligands and death receptor pathway
- •2.2.4. LIF-STAT3 proapoptotic signaling
- •2.2.5. IGF survival signaling
- •2.2.6. Regulation by adhesion
- •2.2.7. PI3K/AKT pathway: molecular hub for survival signals
- •2.2.8. Downstream regulators of apoptosis: the BCL-2 family members
- •3. APOPTOSIS IN BREAST CANCER
- •3.1. Apoptosis in breast tumorigenesis and cancer progression
- •3.2. Molecular dysregulation of apoptosis in breast cancer
- •3.2.1. Altered expression of death ligands and their receptors in breast cancer
- •3.2.2. Deregulation of prosurvival growth factors and their receptors
- •3.2.3. Alterations in cell adhesion and resistance to anoikis
- •3.2.4. Enhanced activation of the PI3K/AKT pathway in breast cancer
- •3.2.5. p53 inactivation in breast cancer
- •3.2.6. Altered expression of BCL-2 family of proteins in breast cancer
- •5. CONCLUSION
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •2. DETECTING CELL DEATH IN THE FEMALE GONADS
- •4. APOPTOSIS AND FEMALE REPRODUCTIVE AGING
- •6. CONCLUDING REMARKS
- •REFERENCES
- •25 Apoptotic Signaling in Male Germ Cells
- •1. INTRODUCTION
- •3.1. Murine models
- •3.2. Primate models
- •3.3. Pathways of caspase activation and apoptosis
- •3.4. Apoptotic signaling in male germ cells
- •5. P38 MITOGEN-ACTIVATED PROTEIN KINASE (MAPK) AND NITRIC OXIDE (NO)–MEDIATED INTRINSIC PATHWAY SIGNALING CONSTITUTES A CRITICAL COMPONENT OF APOPTOTIC SIGNALING IN MALE GERM CELLS AFTER HORMONE DEPRIVATION
- •11. CONCLUSIONS AND PERSPECTIVES
- •REFERENCES
- •26 Cell Death in the Cardiovascular System
- •1. INTRODUCTION
- •2. CELL DEATH IN THE VASCULATURE
- •2.1. Apoptosis in the developing blood vessels
- •2.2. Apoptosis in atherosclerosis
- •2.2.1. Vascular smooth muscle cells
- •2.2.2. Macrophages
- •2.2.3. Regulation of apoptosis in atherosclerosis
- •2.2.4. Necrosis and autophagy in atherosclerosis
- •3. CELL DEATH IN THE MYOCARDIUM
- •3.1. Cell death in myocardial infarction
- •3.1.1. Apoptosis in myocardial infarction
- •3.1.2. Necrosis in myocardial infarction
- •3.1.3. Autophagy in myocardial infarction
- •3.2. Cell death in heart failure
- •3.2.1. Apoptosis in heart failure
- •3.2.2. Necrosis in heart failure
- •3.2.3. Autophagy in heart failure
- •4. CONCLUDING REMARKS
- •ACKNOWLEDGMENTS
- •REFERENCES
- •27 Cell Death Regulation in Muscle
- •1. INTRODUCTION TO MUSCLE
- •1.1. Skeletal muscle adaptation to endurance training
- •1.2. Myonuclear domains
- •2. MITOCHONDRIALLY MEDIATED APOPTOSIS IN MUSCLE
- •2.1. Skeletal muscle apoptotic susceptibility
- •4. APOPTOSIS IN MUSCLE DURING AGING AND DISEASE
- •4.1. Aging
- •4.2. Type 2 diabetes mellitus
- •4.3. Cancer cachexia
- •4.4. Chronic heart failure
- •6. CONCLUSION
- •SUGGESTED READINGS
- •28 Cell Death in the Skin
- •1. INTRODUCTION
- •2. CELL DEATH IN SKIN HOMEOSTASIS
- •2.1. Cornification and apoptosis
- •2.2. Death receptors in the skin
- •3. CELL DEATH IN SKIN PATHOLOGY
- •3.1. Sunburn
- •3.2. Skin cancer
- •3.3. Necrolysis
- •3.4. Pemphigus
- •3.5. Eczema
- •3.6. Graft-versus-host disease
- •4. CONCLUDING REMARKS AND PERSPECTIVES
- •ACKNOWLEDGMENTS
- •SUGGESTED READINGS
- •29 Apoptosis and Cell Survival in the Immune System
- •2.1. Survival of early hematopoietic progenitors
- •2.2. Sizing of the T-cell population
- •2.2.1. Establishing central tolerance
- •2.2.2. Peripheral tolerance
- •2.2.3. Memory T cells
- •2.3. Control of apoptosis in B-cell development
- •2.3.1. Early B-cell development
- •2.3.2. Deletion of autoreactive B cells
- •2.3.3. Survival and death of activated B cells
- •3. IMPAIRED APOPTOSIS AND LEUKEMOGENESIS
- •4. CONCLUSIONS
- •ACKNOWLEDGMENTS
- •REFERENCES
- •30 Cell Death Regulation in the Hematopoietic System
- •1. INTRODUCTION
- •2. HEMATOPOIETIC STEM CELLS
- •4. ERYTHROPOIESIS
- •5. MEGAKARYOPOIESIS
- •6. GRANULOPOIESIS
- •7. MONOPOIESIS
- •8. CONCLUSION
- •ACKNOWLEDGMENTS
- •REFERENCES
- •31 Apoptotic Cell Death in Sepsis
- •1. INTRODUCTION
- •2. HOST INFLAMMATORY RESPONSE TO SEPSIS
- •3. CLINICAL OBSERVATIONS OF CELL DEATH IN SEPSIS
- •3.1. Sepsis-induced apoptosis
- •3.2. Necrotic cell death in sepsis
- •4.1. Central role of apoptosis in sepsis mortality: immune effector cells and gut epithelium
- •4.2. Apoptotic pathways in sepsis-induced immune cell death
- •4.3. Investigations implicating the extrinsic apoptotic pathway in sepsis
- •4.4. Investigations implicating the intrinsic apoptotic pathway in sepsis
- •5. THE EFFECT OF APOPTOSIS ON THE IMMUNE SYSTEM
- •5.1. Cellular effects of an increased apoptotic burdens
- •5.2. Network effects of selective loss of immune cell types
- •5.3. Studies of immunomodulation by apoptotic cells in other fields
- •7. CONCLUSION
- •REFERENCES
- •32 Host–Pathogen Interactions
- •1. INTRODUCTION
- •2. FROM THE PATHOGEN PERSPECTIVE
- •2.1. Commensals versus pathogens
- •2.2. Pathogen strategies to infect the host
- •3. HOST DEFENSE
- •3.1. Antimicrobial peptides
- •3.2. PRRs and inflammation
- •3.2.1. TLRs
- •3.2.2. NLRs
- •3.2.3. The Nod signalosome
- •3.2.4. The inflammasome
- •3.3. Cell death
- •3.3.1. Apoptosis and pathogen clearance
- •3.3.2. Pyroptosis
- •3.2.3. Caspase-independent cell death
- •3.2.4. Autophagy and autophagic cell death
- •4. CONCLUSIONS
- •REFERENCES
- •Part III Cell Death in Nonmammalian Organisms
- •1. PHENOTYPE AND ASSAYS OF YEAST APOPTOSIS
- •2.1. Pheromone-induced cell death
- •2.1.1. Colony growth
- •2.1.2. Killer-induced cell death
- •3. EXTERNAL STIMULI THAT INDUCE APOPTOSIS IN YEAST
- •4. THE GENETICS OF YEAST APOPTOSIS
- •5. PROGRAMMED AND ALTRUISTIC AGING
- •SUGGESTED READINGS
- •34 Caenorhabditis elegans and Apoptosis
- •1. Overview
- •2. KILLING
- •3. SPECIFICATION
- •4. EXECUTION
- •4.1. DNA degradation
- •4.2. Mitochondrial elimination
- •4.3. Engulfment
- •5. SUMMARY
- •SUGGESTED READINGS
- •35 Apoptotic Cell Death in Drosophila
- •2. DROSOPHILA CASPASES AND PROXIMAL REGULATORS
- •6. CLOSING COMMENTS
- •SUGGESTED READINGS
- •36 Analysis of Cell Death in Zebrafish
- •1. INTRODUCTION
- •2. WHY USE ZEBRAFISH TO STUDY CELL DEATH?
- •2.2. Molecular techniques to rapidly assess gene function in embryos
- •2.2.1. Studies of gene function using microinjections into early embryos
- •2.2.2. In situ hybridization and immunohistochemistry
- •2.3. Forward genetic screening
- •2.4. Drug and small-molecule screening
- •2.5. Transgenesis
- •2.6. Targeted knockouts
- •3.1. Intrinsic apoptosis
- •3.2. Extrinsic apoptosis
- •3.3. Chk-1 suppressed apoptosis
- •3.4. Anoikis
- •3.5. Autophagy
- •3.6. Necrosis
- •4. DEVELOPMENTAL CELL DEATH IN ZEBRAFISH EMBRYOS
- •5. THE P53 PATHWAY
- •6. PERSPECTIVES AND FUTURE DIRECTIONS
- •SUGGESTED READING
13 Implications of Nitrosative Stress-Induced Protein
Misfolding in Neurodegeneration
Tomohiro Nakamura and Stuart A. Lipton
SUMMARY
Several chronic neurodegenerative disorders manifest deposits of misfolded or aggregated proteins. Genetic mutations are the root cause for protein misfolding in rare families, but the majority of patients have sporadic forms possibly related to environmental factors. In some cases, the ubiquitin-proteasome system or molecular chaperones can prevent accumulation of aberrantly folded proteins. Recent studies suggest that generation of excessive nitric oxide (NO) and reactive oxygen species, in part due to overactivity of the N-methyl-D-aspartate (NMDA) subtype of glutamate receptor, can mediate protein misfolding in the absence of genetic mutation. S-Nitrosylation, or covalent reaction of NO with specific protein thiol groups, represents one mechanism contributing to NO-induced protein misfolding and neurotoxicity. Here, we present evidence suggesting that NO contributes to protein misfolding via S-nitrosylating protein-disulfide isomerase or the E3 ubiquitin ligase parkin. We discuss how the drugs memantine or NitroMemantine can inhibit excessive NMDA receptor activity to ameliorate NO production, protein misfolding, and neurodegeneration.
1. INTRODUCTION
Many neurodegenerative diseases are characterized by the accumulation of misfolded proteins that adversely affect neuronal connectivity and plasticity and trigger cell death signaling pathways. For example, degenerating brain contains aberrant accumulations of misfolded, aggregated proteins, such as α-synuclein and synphilin- 1 in Parkinson’s disease (PD) and amyloid-β (Aβ) and tau in Alzheimer’s disease (AD). The inclusions observed in PD are called Lewy bodies and are mostly found in the cytoplasm. AD brains show intracellular neurofibrillary tangles, which contain hyperphosphorylated tau, and extracellular plaques, which contain Aβ. These aggregates may consist of oligomeric complexes of nonnative secondary structures and demonstrate poor solubility in aqueous or detergent solvent. Other disorders manifesting protein aggregation include Huntington’s disease
(a polyQ disorder), amyotrophic lateral sclerosis (ALS), and prion disease.
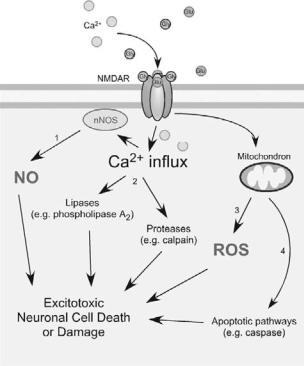
An additional feature of most neurodegenerative diseases is excessive generation of reactive nitrogen species (RNS) and reactive oxygen species (ROS), which can contribute to neuronal cell injury and death. Although many intraand extracellular molecules may participate in neuronal injury, accumulation of nitrosative stress due to excessive generation of nitric oxide (NO) appears to be a potential factor contributing to neuronal cell damage and death. A well-established model for NO production entails a central role of the N-methyl-D- aspartate (NMDA)–type glutamate receptors in the nervous system. Excessive activation of NMDA receptors drives Ca2+ influx, which in turn activates neuronal NO synthase (nNOS), as well as the generation of ROS (Figure 13-1). Accumulating evidence suggests that NO can mediate both protective and neurotoxic effects by reacting with cysteine residues of target proteins to form S- nitrosothiols (SNOs), a process termed S-nitrosylation because of its effects on the chemical biology of protein function. Importantly, normal mitochondrial respiration may also generate free radicals, principally ROS, and one such molecule, superoxide anion (O2.− ), reacts rapidly with free radical NO to form the very toxic product peroxynitrite (ONOO− ).
Importantly, protein aggregation can result from either (1) a rare mutation in the disease-related gene encoding the protein, or (2) post-translational changes to the protein engendered by nitrosative/oxidative stress, which may well account for the more common sporadic cases of the disease. Therefore, a key theme of this article is the hypothesis that nitrosative and oxidative stress contribute to protein misfolding in the brains of the majority of neurodegenerative patients. In this review, we discuss specific examples showing that
145
146 |
TOMOHIRO NAKAMURA AND STUART A. LIPTON |
Figure 13-1. Activation of the NMDA receptor (NMDAR) by glutamate (Glu) and glycine (Gly) induces Ca2+ influx and activates excitotoxic pathways. NMDAR hyperactivation triggers (1) generation of NO, (2) activation of lipases and proteases, (3) production and release of ROS from mitochondria, and (4) activation of caspases, contributing to neuronal cell death and damage.
S-nitrosylation of (1) ubiquitin E3 ligases such as parkin or (2) endoplasmic reticulum chaperones such as protein-disulfide isomerase (PDI) is critical for the accumulation of misfolded proteins in neurodegenerative diseases such as PD and other conditions. We also discuss the neuroprotective mechanism of action of NMDA open-channel blockers like memantine and NO-related drugs for the treatment of neurodegenerative disorders.
2. PROTEIN MISFOLDING AND AGGREGATION IN
NEURODEGENERATIVE DISEASES
In general, protein aggregates do not accumulate in unstressed, healthy neurons due in part to the existence of cellular “quality control machineries.” For example, molecular chaperones are believed to provide a defense mechanism against the toxicity of misfolded proteins because chaperones can prevent inappropriate interactions within and between polypeptides and can promote refolding of proteins that have been misfolded because of cell stress. In addition to the quality control of proteins provided by molecular chaperones, the ubiquitin– proteasome system (UPS) and autophagy/lysosomal degradation are involved in the clearance of abnormal
or aberrant proteins. When chaperones cannot repair misfolded proteins, they may be tagged via addition of polyubiquitin chains for degradation by the proteasome. In neurodegenerative conditions, intraor extracellular protein aggregates are thought to accumulate in the brain as a result of a decrease in molecular chaperone or proteasome activities. In fact, several mutations that disturb the activity of molecular chaperones or UPS-associated enzymes can cause neurodegeneration. Along these lines, postmortem samples from the substantia nigra of PD patients (vs. non-PD controls) manifest a significant reduction in proteasome activity.
Historically, lesions that contain aggregated proteins were considered to be pathogenic. Recently, several lines of evidence have suggested that aggregates are formed through a complex multistep process by which misfolded proteins assemble into inclusion bodies; currently, soluble (micro-)oligomers of these aberrant proteins are thought to be the most toxic forms via interference with normal cell activities, whereas frank macroscopic aggregates may be an attempt by the cell to wall off potentially toxic material.
3. NMDA RECEPTOR-MEDIATED GLUTAMATERGIC SIGNALING PATHWAYS INDUCE CA2+ INFLUX AND
GENERATION OF RNS/ROS
It is well known that the amino acid glutamate is the major excitatory neurotransmitter in the brain. Glutamate is present in high concentrations in the adult central nervous system and is released for milliseconds from nerve terminals in a Ca2+ -dependent manner. After glutamate enters the synaptic cleft, it diffuses across the cleft to interact with its corresponding receptors on the postsynaptic face of an adjacent neuron. Excitatory neurotransmission is necessary for the normal development and plasticity of synapses and for some forms of learning or memory; however, excessive activation of glutamate receptors is implicated in neuronal damage in many neurological disorders, ranging from acute hypoxic-ischemic brain injury to chronic neurodegenerative diseases. John Olney coined the term excitotoxicity to describe the toxic effect of glutamate. It is currently thought that overstimulation of extrasynaptic NMDA receptors mediate, at least in part, this type of neuronal damage, whereas, in contrast, synaptic activity predominantly activates survival pathways. Intense hyperstimulation of excitatory receptors leads to necrotic cell death, but more mild or chronic overstimulation can result in apoptotic or other forms of cell death.
There are two large families of glutamate receptors in the nervous system, ionotropic receptors (representing
IMPLICATIONS OF NITROSATIVE STRESS-INDUCED PROTEIN MISFOLDING IN NEURODEGENERATION |
147 |
ligand-gated ion channels) and metabotropic receptors (coupled to G-proteins). Ionotropic glutamate receptors are further divided into three broad classes: (1) NMDA receptors, (2) α-amino–3-hydroxy-5 methyl-4-isoxazole propionic acid (AMPA) receptors, and (3) kainate receptors, which are each named after synthetic ligands that can selectively activate these receptors. The NMDA receptor has attracted attention for a long period of time because it has several properties that set it apart from other ionotropic glutamate receptors. One such characteristic, in contrast to most AMPA and kainate receptors, is that NMDA receptor-coupled channels are highly permeable to Ca2+, thus permitting Ca2+ entry after ligand binding if the cell is depolarized to relieve block of the receptor-associated ion channel by Mg2+. Subsequent binding of Ca2+ to various intracellular molecules can lead to many significant consequences. For instance, increased levels of neuronal Ca2+, in conjunction with the Ca2+-binding protein CaM, trigger the activation of nNOS and subsequent generation of NO from the amino acid L-arginine. NO is a gaseous free radical (thus highly diffusible) and a key molecule that plays a vital role in normal signal transduction, but in excess it can lead to neuronal cell damage and death. In particular, excessive activation of NMDA receptors leads to the production of damaging free radicals (e.g., NO and ROS) and other enzymatic processes, contributing to cell death (Figure 13-1).
Increased nitrosative and oxidative stress are associated with chaperone and proteasomal dysfunction, resulting in accumulation of misfolded aggregates. However, until recently, little was known regarding the molecular and pathogenic mechanisms underlying contribution of NO to the formation of inclusion bodies such as amyloid plaques in AD or Lewy bodies in PD.
4. PROTEIN S-NITROSYLATION AND NEURONAL
CELL DEATH
Early investigations indicated that NO participates in cellular signaling pathways, which regulate broad aspects of brain function, including synaptic plasticity, normal development, and neuronal cell death. In general, NO exerts physiologic and some pathophysiologic effects via stimulation of guanylate cyclase to form cyclic guanosine-3 ,5 -monophosphate or through S-nitros(yl)ation of regulatory protein thiol groups. S- Nitrosylation is the covalent addition of an NO group to a critical cysteine thiol/sulfhydryl (or, more properly, thiolate anion, RS−) to form an S-nitrosothiol derivative (R-SNO). Such modification modulates the function of a broad spectrum of mammalian, plant, and microbial
proteins. In general, a consensus motif of amino acids comprised of nucleophilic residues (generally an acid and a base) surround a critical cysteine, which increases the cysteine sulfhydryl’s susceptibility to S-nitrosylation. Our group first identified the physiologic relevance of S- nitrosylation by showing that NO and related RNS exert paradoxical effects via redox-based mechanisms – NO is neuroprotective via S-nitrosylation of NMDA receptors (as well as other subsequently discovered targets, including caspases) and yet can also be neurodestructive by formation of peroxynitrite (or, as later discovered, reaction with additional molecules such as matrix metalloproteinase 9 and glyceraldehyde 3-phosphate dehydrogenase [GAPDH]). Over the past decade, accumulating evidence has suggested that S-nitrosylation can regulate the biological activity of a great variety of proteins, in some ways akin to phosphorylation. Chemically, NO is often a good “leaving group,” facilitating further oxidation of critical thiol to disulfide bonds among neighboring (vicinal) cysteine residues or, via reaction with ROS, to sulfenic (-SOH), sulfinic (-SO2H) or sulfonic (-SO3H) acid derivitization of the protein.
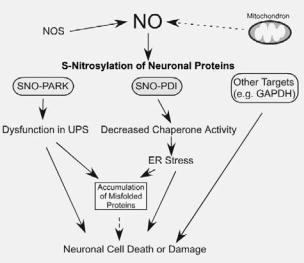
Inhibition of NOS activity ameliorates the progression of disease pathology in animal models of PD, AD, and ALS, suggesting that excess generation of NO plays a pivotal role in the pathogenesis of several neurodegenerative diseases. Although the involvement of NO in neurodegeneration has been widely accepted, the chemical relationship between nitrosative stress and accumulation of misfolded proteins has remained obscure. Recent findings, however, have shed light on molecular events underlying this relationship. Specifically, we recently mounted physiologic and chemical evidence that S-nitrosylation modulates the (1) ubiquitin E3 ligase activity of parkin, and (2) chaperone and isomerase activities of PDI, contributing to protein misfolding and neurotoxicity in models of neurodegenerative disorders (Figure 13-2).
5. S-NITROSYLATION OF PARKIN
Identification of errors in the genes encoding parkin (a ubiquitin E3 ligase) and UCH-L1 (deubiquitinating enzyme) in rare familial forms of PD has implicated possible dysfunction of the UPS in the pathogenesis of sporadic PD. The UPS represents an important mechanism for proteolysis in mammalian cells. Formation of polyubiquitin chains constitutes the signal for proteasomal attack and degradation. An isopeptide bond covalently attaches the C-terminus of the first ubiquitin in a polyubiquitin chain to a lysine residue in the target protein. The cascade of activating (E1), conjugating (E2),
148 |
TOMOHIRO NAKAMURA AND STUART A. LIPTON |
Figure 13-2. Possible mechanism whereby S-nitrosylated species contribute to the accumulation of aberrant proteins and neuronal damage. S-nitrosylation of parkin (forming SNO-PARK) and PDI (forming SNO-PDI) can contribute to neuronal cell injury in part by triggering accumulation of misfolded proteins. S-Nitrosylation of other proteins, such as glyceraldehyde 3-phosphate dehydrogenase (GAPDH), may also contribute to neuronal cell injury or death.
and ubiquitin-ligating (E3) type enzymes catalyzes the conjugation of the ubiquitin chain to proteins. In addition, individual E3 ubiquitin ligases play a key role in the recognition of specific substrates.
Mutations in the parkin gene can cause autosomalrecessive juvenile Parkinsonism (ARJP), accounting for some cases of hereditary PD manifest in young patients with onset beginning anywhere from the teenage years through the 40s. Parkin is a member of a large family of E3 ubiquitin ligases that are related to one another by the presence of RING (really interesting new gene) finger domains. Point mutations, stop mutations, truncations, and deletions in both alleles of the parkin gene will eventually cause dysfunction in its activity and are responsible for many cases of ARJP as well as rare adult forms of PD.
Nitrosative/oxidative stress, commonly found during normal aging, can mimic rare genetic causes of disorders, such as PD, by promoting protein misfolding in the absence of a genetic mutation. For example, S-nitrosylation and further oxidation of parkin or UchL1 result in dysfunction of these enzymes and thus of the UPS (Figure 13-2). We and others recently discovered that nitrosative stress triggers S-nitrosylation of parkin (forming SNO-parkin) not only in rodent models of PD, but also in the brains of human patients with PD and the related α-synucleinopathy, diffuse Lewy body disease. SNO-parkin initially stimulates ubiquitin E3 ligase activity, resulting in enhanced ubiquitination as observed in
Lewy bodies, followed by a decrease in enzyme activity, producing a futile cycle of dysfunctional UPS (Figure 13-2). We also found that rotenone led to the generation of SNO-parkin and thus dysfunctional ubiquitin E3 ligase activity. Moreover, S-nitrosylation appears to compromise the neuroprotective effect of parkin. These mechanisms involve S-nitrosylation of critical cysteine residues in the first RING domain of parkin. Nitrosative and oxidative stress can also alter the solubility of parkin via post-translational modification of cysteine residues, which may concomitantly compromise its protective function.
6. S-NITROSYLATION OF PDI MEDIATES PROTEIN
MISFOLDING AND NEUROTOXICITY IN CELL MODELS
OF PD OR AD
The endoplasmic reticulum (ER) normally participates in protein processing and folding but undergoes a stress response when immature or misfolded proteins accumulate. ER stress stimulates two critical intracellular responses. The first represents expression of chaperones that prevent protein aggregation via the unfolded protein response (UPR) and is implicated in protein refolding, post-translational assembly of protein complexes, and protein degradation. The second ER stress response, termed ER-associated degradation, specifically recognizes terminally misfolded proteins for retrotranslocation across the ER membrane to the cytosol, where they can be degraded by the UPS. Additionally, although severe ER stress or a prolonged UPR can induce apoptosis, the ER withstands relatively mild insults via expression of stress proteins such as glucose-regulated protein and PDI. These proteins behave as molecular chaperones that assist in the maturation, transport, and folding of secretory proteins.
During protein folding in the ER, PDI can also introduce disulfide bonds into proteins (oxidation), break disulfide bonds (reduction), and catalyze thiol/disulfide exchange (isomerization), thus facilitating disulfide bond formation, rearrangement reactions, and structural stability. PDI has two redox active CXXC motifs, and these two-thiol/disulfide centers function as independent active sites.
In many neurodegenerative disorders and cerebral ischemia, the accumulation of immature and denatured proteins results in ER dysfunction, but upregulation of PDI represents an adaptive response promoting protein refolding and may offer neuronal cell protection. In addition, it is generally accepted that excessive generation of NO can contribute to activation of the ER stress pathway, at least in some cell types. Molecular mechanisms
