
- •Contents
- •Contributors
- •Part I General Principles of Cell Death
- •1 Human Caspases – Apoptosis and Inflammation Signaling Proteases
- •1.1. Apoptosis and limited proteolysis
- •1.2. Caspase evolution
- •2. ACTIVATION MECHANISMS
- •2.2. The activation platforms
- •2.4. Proteolytic maturation
- •3. CASPASE SUBSTRATES
- •4. REGULATION BY NATURAL INHIBITORS
- •REFERENCES
- •2 Inhibitor of Apoptosis Proteins
- •2. CELLULAR FUNCTIONS AND PHENOTYPES OF IAP
- •3. IN VIVO FUNCTIONS OF IAP FAMILY PROTEINS
- •4. SUBCELLULAR LOCATIONS OF IAP
- •8. IAP–IAP INTERACTIONS
- •10. ENDOGENOUS ANTAGONISTS OF IAP
- •11. IAPs AND DISEASE
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •2.1. The CD95 (Fas/APO-1) system
- •2.1.1. CD95 and CD95L: discovery of the first direct apoptosis-inducing receptor-ligand system
- •2.1.2. Biochemistry of CD95 apoptosis signaling
- •2.2. The TRAIL (Apo2L) system
- •3.1. The TNF system
- •3.1.1. Biochemistry of TNF signal transduction
- •3.1.2. TNF and TNF blockers in the clinic
- •3.2. The DR3 system
- •4. THE DR6 SYSTEM
- •6. CONCLUDING REMARKS AND OUTLOOK
- •SUGGESTED READINGS
- •4 Mitochondria and Cell Death
- •1. INTRODUCTION
- •2. MITOCHONDRIAL PHYSIOLOGY
- •3. THE MITOCHONDRIAL PATHWAY OF APOPTOSIS
- •9. CONCLUSIONS
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •3. INHIBITING APOPTOSIS
- •4. INHIBITING THE INHIBITORS
- •6. THE BCL-2 FAMILY AND CANCER
- •SUGGESTED READINGS
- •6 Endoplasmic Reticulum Stress Response in Cell Death and Cell Survival
- •1. INTRODUCTION
- •2. THE ESR IN YEAST
- •3. THE ESR IN MAMMALS
- •4. THE ESR AND CELL DEATH
- •5. THE ESR IN DEVELOPMENT AND TISSUE HOMEOSTASIS
- •6. THE ESR IN HUMAN DISEASE
- •7. CONCLUSION
- •7 Autophagy – The Liaison between the Lysosomal System and Cell Death
- •1. INTRODUCTION
- •2. AUTOPHAGY
- •2.2. Physiologic functions of autophagy
- •2.3. Autophagy and human pathology
- •3. AUTOPHAGY AND CELL DEATH
- •3.1. Autophagy as anti–cell death mechanism
- •3.2. Autophagy as a cell death mechanism
- •3.3. Molecular players of the autophagy–cell death cross-talk
- •4. AUTOPHAGY, CELLULAR DEATH, AND CANCER
- •5. CONCLUDING REMARKS AND PENDING QUESTIONS
- •SUGGESTED READINGS
- •8 Cell Death in Response to Genotoxic Stress and DNA Damage
- •1. TYPES OF DNA DAMAGE AND REPAIR SYSTEMS
- •2. DNA DAMAGE RESPONSE
- •2.2. Transducers
- •2.3. Effectors
- •4. CHROMATIN MODIFICATIONS
- •5. CELL CYCLE CHECKPOINT REGULATION
- •6. WHEN REPAIR FAILS: SENESCENCE VERSUS APOPTOSIS
- •6.1. DNA damage response and the induction of apoptosis
- •6.2. p53-independent mechanisms of apoptosis
- •6.3. DNA damage response and senescence induction
- •7. DNA DAMAGE FROM OXIDATIVE STRESS
- •SUGGESTED READINGS
- •9 Ceramide and Lipid Mediators in Apoptosis
- •1. INTRODUCTION
- •3.1. Basic cell signaling often involves small molecules
- •3.2. Sphingolipids are cell-signaling molecules
- •3.2.1. Ceramide induces apoptosis
- •3.2.2. Ceramide accumulates during programmed cell death
- •3.2.3. Inhibition of ceramide production alters cell death signaling
- •4.1. Ceramide is generated through SM hydrolysis
- •4.3. aSMase can be activated independently of extracellular receptors to regulate apoptosis
- •4.4. Controversial aspects of the role of aSMase in apoptosis
- •4.5. De novo ceramide synthesis regulates programmed cell death
- •4.6. p53 and Bcl-2–like proteins are connected to de novo ceramide synthesis
- •4.7. The role and regulation of de novo synthesis in ceramide-mediated cell death is poorly understood
- •5. CONCLUDING REMARKS AND FUTURE DIRECTIONS
- •5.1. Who? (Which enzyme?)
- •5.2. What? (Which ceramide?)
- •5.3. Where? (Which compartment?)
- •5.4. When? (At what steps?)
- •5.5. How? (Through what mechanisms?)
- •5.6. What purpose?
- •6. SUMMARY
- •SUGGESTED READINGS
- •1. General Introduction
- •1.1. Cytotoxic lymphocytes and apoptosis
- •2. CYTOTOXIC GRANULES AND GRANULE EXOCYTOSIS
- •2.1. Synthesis and loading of the cytotoxic granule proteins into the secretory granules
- •2.2. The immunological synapse
- •2.3. Secretion of granule proteins
- •2.4. Uptake of proapoptotic proteins into the target cell
- •2.5. Activation of death pathways by granzymes
- •3. GRANULE-BOUND CYTOTOXIC PROTEINS
- •3.1. Perforin
- •3.2. Granulysin
- •3.3. Granzymes
- •3.3.1. GrB-mediated apoptosis
- •3.3.2. GrA-mediated cell death
- •3.3.3. Orphan granzyme-mediated cell death
- •5. CONCLUSIONS
- •REFERENCES
- •Part II Cell Death in Tissues and Organs
- •1.1. Death by trophic factor deprivation
- •1.2. Key molecules regulating neuronal apoptosis during development
- •1.2.1. Roles of caspases and Apaf-1 in neuronal cell death
- •1.2.2. Role of Bcl-2 family members in neuronal cell death
- •1.3. Signal transduction from neurotrophins and neurotrophin receptors
- •1.3.1. Signals for survival
- •1.3.2. Signals for death
- •2.1. Apoptosis in neurodegenerative diseases
- •2.1.4. Amyotrophic lateral sclerosis
- •2.2. Necrotic cell death in neurodegenerative diseases
- •2.2.1. Calpains
- •2.2.2. Cathepsins
- •3. CONCLUSIONS
- •ACKNOWLEDGMENT
- •SUGGESTED READINGS
- •ACKNOWLEDGMENT
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •5. S-NITROSYLATION OF PARKIN
- •7. POTENTIAL TREATMENT OF EXCESSIVE NMDA-INDUCED Ca2+ INFLUX AND FREE RADICAL GENERATION
- •8. FUTURE THERAPEUTICS: NITROMEMANTINES
- •9. CONCLUSIONS
- •Acknowledgments
- •SUGGESTED READINGS
- •3. MITOCHONDRIAL PERMEABILITY TRANSITION ACTIVATED BY Ca2+ AND OXIDATIVE STRESS
- •4.1. Mitochondrial apoptotic pathways
- •4.2. Bcl-2 family proteins
- •4.3. Caspase-dependent apoptosis
- •4.4. Caspase-independent apoptosis
- •4.5. Calpains in ischemic neural cell death
- •5. SUMMARY
- •ACKNOWLEDGMENTS
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •2. HISTORICAL ANTECEDENTS
- •7.1. Activation of p21 waf1/cip1: Targeting extrinsic and intrinsic pathways to death
- •8. CONCLUSION
- •ACKNOWLEDGMENTS
- •REFERENCES
- •16 Apoptosis and Homeostasis in the Eye
- •1.1. Lens
- •1.2. Retina
- •2. ROLE OF APOPTOSIS IN DISEASES OF THE EYE
- •2.1. Glaucoma
- •2.2. Age-related macular degeneration
- •4. APOPTOSIS AND OCULAR IMMUNE PRIVILEGE
- •5. CONCLUSIONS
- •SUGGESTED READINGS
- •17 Cell Death in the Inner Ear
- •3. THE COCHLEA IS THE HEARING ORGAN
- •3.1. Ototoxic hair cell death
- •3.2. Aminoglycoside-induced hair cell death
- •3.3. Cisplatin-induced hair cell death
- •3.4. Therapeutic strategies to prevent hair cell death
- •3.5. Challenges to studies of hair cell death
- •4. SPIRAL GANGLION NEURON DEATH
- •4.1. Neurotrophic support from sensory hair cells and supporting cells
- •4.2. Afferent activity from hair cells
- •4.3. Molecular manifestations of spiral ganglion neuron death
- •4.4. Therapeutic interventions to prevent SGN death
- •ACKNOWLEDGMENTS
- •SUGGESTED READINGS
- •18 Cell Death in the Olfactory System
- •1. Introduction
- •2. Anatomical Aspects
- •3. Life and Death in the Olfactory System
- •3.1. Olfactory epithelium
- •3.2. Olfactory bulb
- •REFERENCES
- •1. Introduction
- •3.1. Beta cell death in the development of T1D
- •3.2. Mechanisms of beta cell death in type 1 diabetes
- •3.2.1. Apoptosis signaling pathways downstream of death receptors and inflammatory cytokines
- •3.2.2. Oxidative stress
- •3.3. Mechanisms of beta cell death in type 2 diabetes
- •3.3.1. Glucolipitoxicity
- •3.3.2. Endoplasmic reticulum stress
- •5. SUMMARY
- •Acknowledgments
- •REFERENCES
- •20 Apoptosis in the Physiology and Diseases of the Respiratory Tract
- •1. APOPTOSIS IN LUNG DEVELOPMENT
- •2. APOPTOSIS IN LUNG PATHOPHYSIOLOGY
- •2.1. Apoptosis in pulmonary inflammation
- •2.2. Apoptosis in acute lung injury
- •2.3. Apoptosis in chronic obstructive pulmonary disease
- •2.4. Apoptosis in interstitial lung diseases
- •2.5. Apoptosis in pulmonary arterial hypertension
- •2.6. Apoptosis in lung cancer
- •SUGGESTED READINGS
- •21 Regulation of Cell Death in the Gastrointestinal Tract
- •1. INTRODUCTION
- •2. ESOPHAGUS
- •3. STOMACH
- •4. SMALL AND LARGE INTESTINE
- •5. LIVER
- •6. PANCREAS
- •7. SUMMARY AND CONCLUDING REMARKS
- •SUGGESTED READINGS
- •22 Apoptosis in the Kidney
- •1. NORMAL KIDNEY STRUCTURE AND FUNCTION
- •3. APOPTOSIS IN ADULT KIDNEY DISEASE
- •4. REGULATION OF APOPTOSIS IN KIDNEY CELLS
- •4.1. Survival factors
- •4.2. Lethal factors
- •4.2.1. TNF superfamily cytokines
- •4.2.2. Other cytokines
- •4.2.3. Glucose
- •4.2.4. Drugs and xenobiotics
- •4.2.5. Ischemia-reperfusion and sepsis
- •5. THERAPEUTIC APPROACHES
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •2. APOPTOSIS IN THE NORMAL BREAST
- •2.1. Occurrence and role of apoptosis in the developing breast
- •2.2.2. Death ligands and death receptor pathway
- •2.2.4. LIF-STAT3 proapoptotic signaling
- •2.2.5. IGF survival signaling
- •2.2.6. Regulation by adhesion
- •2.2.7. PI3K/AKT pathway: molecular hub for survival signals
- •2.2.8. Downstream regulators of apoptosis: the BCL-2 family members
- •3. APOPTOSIS IN BREAST CANCER
- •3.1. Apoptosis in breast tumorigenesis and cancer progression
- •3.2. Molecular dysregulation of apoptosis in breast cancer
- •3.2.1. Altered expression of death ligands and their receptors in breast cancer
- •3.2.2. Deregulation of prosurvival growth factors and their receptors
- •3.2.3. Alterations in cell adhesion and resistance to anoikis
- •3.2.4. Enhanced activation of the PI3K/AKT pathway in breast cancer
- •3.2.5. p53 inactivation in breast cancer
- •3.2.6. Altered expression of BCL-2 family of proteins in breast cancer
- •5. CONCLUSION
- •SUGGESTED READINGS
- •1. INTRODUCTION
- •2. DETECTING CELL DEATH IN THE FEMALE GONADS
- •4. APOPTOSIS AND FEMALE REPRODUCTIVE AGING
- •6. CONCLUDING REMARKS
- •REFERENCES
- •25 Apoptotic Signaling in Male Germ Cells
- •1. INTRODUCTION
- •3.1. Murine models
- •3.2. Primate models
- •3.3. Pathways of caspase activation and apoptosis
- •3.4. Apoptotic signaling in male germ cells
- •5. P38 MITOGEN-ACTIVATED PROTEIN KINASE (MAPK) AND NITRIC OXIDE (NO)–MEDIATED INTRINSIC PATHWAY SIGNALING CONSTITUTES A CRITICAL COMPONENT OF APOPTOTIC SIGNALING IN MALE GERM CELLS AFTER HORMONE DEPRIVATION
- •11. CONCLUSIONS AND PERSPECTIVES
- •REFERENCES
- •26 Cell Death in the Cardiovascular System
- •1. INTRODUCTION
- •2. CELL DEATH IN THE VASCULATURE
- •2.1. Apoptosis in the developing blood vessels
- •2.2. Apoptosis in atherosclerosis
- •2.2.1. Vascular smooth muscle cells
- •2.2.2. Macrophages
- •2.2.3. Regulation of apoptosis in atherosclerosis
- •2.2.4. Necrosis and autophagy in atherosclerosis
- •3. CELL DEATH IN THE MYOCARDIUM
- •3.1. Cell death in myocardial infarction
- •3.1.1. Apoptosis in myocardial infarction
- •3.1.2. Necrosis in myocardial infarction
- •3.1.3. Autophagy in myocardial infarction
- •3.2. Cell death in heart failure
- •3.2.1. Apoptosis in heart failure
- •3.2.2. Necrosis in heart failure
- •3.2.3. Autophagy in heart failure
- •4. CONCLUDING REMARKS
- •ACKNOWLEDGMENTS
- •REFERENCES
- •27 Cell Death Regulation in Muscle
- •1. INTRODUCTION TO MUSCLE
- •1.1. Skeletal muscle adaptation to endurance training
- •1.2. Myonuclear domains
- •2. MITOCHONDRIALLY MEDIATED APOPTOSIS IN MUSCLE
- •2.1. Skeletal muscle apoptotic susceptibility
- •4. APOPTOSIS IN MUSCLE DURING AGING AND DISEASE
- •4.1. Aging
- •4.2. Type 2 diabetes mellitus
- •4.3. Cancer cachexia
- •4.4. Chronic heart failure
- •6. CONCLUSION
- •SUGGESTED READINGS
- •28 Cell Death in the Skin
- •1. INTRODUCTION
- •2. CELL DEATH IN SKIN HOMEOSTASIS
- •2.1. Cornification and apoptosis
- •2.2. Death receptors in the skin
- •3. CELL DEATH IN SKIN PATHOLOGY
- •3.1. Sunburn
- •3.2. Skin cancer
- •3.3. Necrolysis
- •3.4. Pemphigus
- •3.5. Eczema
- •3.6. Graft-versus-host disease
- •4. CONCLUDING REMARKS AND PERSPECTIVES
- •ACKNOWLEDGMENTS
- •SUGGESTED READINGS
- •29 Apoptosis and Cell Survival in the Immune System
- •2.1. Survival of early hematopoietic progenitors
- •2.2. Sizing of the T-cell population
- •2.2.1. Establishing central tolerance
- •2.2.2. Peripheral tolerance
- •2.2.3. Memory T cells
- •2.3. Control of apoptosis in B-cell development
- •2.3.1. Early B-cell development
- •2.3.2. Deletion of autoreactive B cells
- •2.3.3. Survival and death of activated B cells
- •3. IMPAIRED APOPTOSIS AND LEUKEMOGENESIS
- •4. CONCLUSIONS
- •ACKNOWLEDGMENTS
- •REFERENCES
- •30 Cell Death Regulation in the Hematopoietic System
- •1. INTRODUCTION
- •2. HEMATOPOIETIC STEM CELLS
- •4. ERYTHROPOIESIS
- •5. MEGAKARYOPOIESIS
- •6. GRANULOPOIESIS
- •7. MONOPOIESIS
- •8. CONCLUSION
- •ACKNOWLEDGMENTS
- •REFERENCES
- •31 Apoptotic Cell Death in Sepsis
- •1. INTRODUCTION
- •2. HOST INFLAMMATORY RESPONSE TO SEPSIS
- •3. CLINICAL OBSERVATIONS OF CELL DEATH IN SEPSIS
- •3.1. Sepsis-induced apoptosis
- •3.2. Necrotic cell death in sepsis
- •4.1. Central role of apoptosis in sepsis mortality: immune effector cells and gut epithelium
- •4.2. Apoptotic pathways in sepsis-induced immune cell death
- •4.3. Investigations implicating the extrinsic apoptotic pathway in sepsis
- •4.4. Investigations implicating the intrinsic apoptotic pathway in sepsis
- •5. THE EFFECT OF APOPTOSIS ON THE IMMUNE SYSTEM
- •5.1. Cellular effects of an increased apoptotic burdens
- •5.2. Network effects of selective loss of immune cell types
- •5.3. Studies of immunomodulation by apoptotic cells in other fields
- •7. CONCLUSION
- •REFERENCES
- •32 Host–Pathogen Interactions
- •1. INTRODUCTION
- •2. FROM THE PATHOGEN PERSPECTIVE
- •2.1. Commensals versus pathogens
- •2.2. Pathogen strategies to infect the host
- •3. HOST DEFENSE
- •3.1. Antimicrobial peptides
- •3.2. PRRs and inflammation
- •3.2.1. TLRs
- •3.2.2. NLRs
- •3.2.3. The Nod signalosome
- •3.2.4. The inflammasome
- •3.3. Cell death
- •3.3.1. Apoptosis and pathogen clearance
- •3.3.2. Pyroptosis
- •3.2.3. Caspase-independent cell death
- •3.2.4. Autophagy and autophagic cell death
- •4. CONCLUSIONS
- •REFERENCES
- •Part III Cell Death in Nonmammalian Organisms
- •1. PHENOTYPE AND ASSAYS OF YEAST APOPTOSIS
- •2.1. Pheromone-induced cell death
- •2.1.1. Colony growth
- •2.1.2. Killer-induced cell death
- •3. EXTERNAL STIMULI THAT INDUCE APOPTOSIS IN YEAST
- •4. THE GENETICS OF YEAST APOPTOSIS
- •5. PROGRAMMED AND ALTRUISTIC AGING
- •SUGGESTED READINGS
- •34 Caenorhabditis elegans and Apoptosis
- •1. Overview
- •2. KILLING
- •3. SPECIFICATION
- •4. EXECUTION
- •4.1. DNA degradation
- •4.2. Mitochondrial elimination
- •4.3. Engulfment
- •5. SUMMARY
- •SUGGESTED READINGS
- •35 Apoptotic Cell Death in Drosophila
- •2. DROSOPHILA CASPASES AND PROXIMAL REGULATORS
- •6. CLOSING COMMENTS
- •SUGGESTED READINGS
- •36 Analysis of Cell Death in Zebrafish
- •1. INTRODUCTION
- •2. WHY USE ZEBRAFISH TO STUDY CELL DEATH?
- •2.2. Molecular techniques to rapidly assess gene function in embryos
- •2.2.1. Studies of gene function using microinjections into early embryos
- •2.2.2. In situ hybridization and immunohistochemistry
- •2.3. Forward genetic screening
- •2.4. Drug and small-molecule screening
- •2.5. Transgenesis
- •2.6. Targeted knockouts
- •3.1. Intrinsic apoptosis
- •3.2. Extrinsic apoptosis
- •3.3. Chk-1 suppressed apoptosis
- •3.4. Anoikis
- •3.5. Autophagy
- •3.6. Necrosis
- •4. DEVELOPMENTAL CELL DEATH IN ZEBRAFISH EMBRYOS
- •5. THE P53 PATHWAY
- •6. PERSPECTIVES AND FUTURE DIRECTIONS
- •SUGGESTED READING
30 |
HENNING WALCZAK AND CHAHRAZADE KANTARI |
for the best possible clinical use of the different TRAIL receptor agonists, it is of pivotal importance that we spend more time and effort on the thorough understanding of the biochemical mechanisms of TRAIL apoptosis resistance versus sensitivity of different types of cancer cells that rely on specific combinations of alterations in the expression of a particular set of oncogenes and tumor suppressor genes as compared with normal cells. Thereby we may be able in the future to specifically target expression or activity of certain factors to achieve cancer-specific TRAIL apoptosis sensitization on the level of the individual cancer patient. Novel proteomic and genomic technologies and the integration of the results obtained by their application in intelligent systems biology approaches will most likely be instrumental in uncovering the mechanisms that govern TRAIL apoptosis sensitivity versus resistance in different types of cancer. These studies will enable a more targeted and individualized use of TRAIL receptor agonists and combination with other drugs in cancer therapy in the future.
3. DEATH RECEPTOR–LIGAND SYSTEMS WITH PRIMARILY
IMMUNOSTIMULATORY, PROINFLAMMATORY ACTIVITY
3.1. The TNF system
3.1.1. Biochemistry of TNF signal transduction
The founding member of the TNFSF is a homotrimer of TNF molecules, each 157 amino acids in length. The trimer adopts a characteristic conformation, which is now commonly referred to as the TNF fold. TNF is mainly produced by activated macrophages. Depending on the physiologic or pathological context, it is, however, also expressed by a number of other cell types. The binding of TNF to its receptors triggers a series of intracellular events that primarily induces the activation of NF-κB and the mitogen-activated protein (MAP) kinases c-Jun N-terminal kinase (JNK) and p38. These events lead to immunostimulatory gene induction, which often drives an inflammatory response (Figure 3-3). Induction of apoptosis by TNF is only a secondary signal (see Section 5).
The diverse biological effects of TNF are mediated by two different receptors, TNF-R1 and TNF-R2. Although TNF-R1 is expressed on cells of almost all tissues, TNFR2 is almost exclusively present on cells of lymphoid origin. TNF-R1 contains a DD and initiates the majority of TNF-induced biological activities, including induction of cell death by apoptosis. Yet TNF-R2 was also shown to be capable of inducing apoptosis. It has now been demonstrated, however, that TNF-R2–induced apopto-
sis works via an indirect loop mechanism: TNF-R2 crosslinking induces expression of TNF, which then binds to TNF-R1 to induce cell death. Apart from inducing TNF, the TNF-R2–mediated signal also sensitizes cells to TNF-R1–mediated apoptosis by depleting TRAF2 and cIAPs, which causes the gene-inducing capacity of TNFR1 to be diminished, strengthening the apoptotic arm of the response (Figure 3-4). Thus the TNF-R2 signal is a modulator of the TNF-R1 signal transduction machinery, and other non–DD-containing receptors of the TNFRSF described to induce apoptosis in certain cells, including CD40, CD30, and FN14, also induce TNF and therefore work in a fashion similar to TNF-R2. Because TNF-R1 is the main signaling receptor for TNF, we now examine its activities in more detail. Binding of TNF to TNFR1 induces receptor oligomerization and recruitment of cytoplasmic signaling proteins, leading to the formation of the TNF-R1 signaling complex (TNF-RSC). The composition of the TNF-RSC and the following steps in TNF-R1–mediated signaling have been extensively studied over the last decade. Although further analysis will be required to discover all the players involved in this process, it is fair to say that to date, it is one of the best understood receptor signaling complexes and cascades in cell biology.
On activation of TNF-R1 by TNF-induced crosslinking at the plasma membrane, the TNFR1 DD serves as a docking site for the DD-containing adaptor protein TRADD. TRADD is recruited to the DD of TNF-R1 via a homotypic DD interaction. TRADD in turn recruits the TNF-R–associated factor-2 (TRAF2) and the receptorinteracting protein 1 (RIP1), a serine/threonine kinase. RIP1, however, can also directly bind to TNF-R1 via its own DD without the need for TRADD. The importance of this interaction remains unclear. Recruitment of TRAF2 (or TRAF5) by TRADD enables recruitment of cIAP1 and/or cIAP2 to the TNF-RSC. The ubiquitin ligase activities of both TRAFs and cIAPs are required for decoration of RIP1 by polyubiquitin chains and for NF-κB and MAP kinase activation. Ubiquitin chains can be formed via linkages of the ubiquitin subunits on different ε-amino groups of the seven different lysines present in ubiquitin or via the α-amino group at the amino-terminus of ubiquitin, with the latter creating linear ubiquitin chains. Thus far it is thought that polyubiquitin chains involved in TNF signaling are either linked via the ε-amino groups of lysine 63 (K63) or K48 of ubiquitin. However, recent data obtained by us and others revealed that linear ubiquitin chains also play an important role in this process. A protein complex termed LUBAC (for linear ubiquitin chain assembly complex) forms an integral part of the TNF-R1 signaling complex (Haas et al., 2009). Furthermore, LUBAC is required for

DEATH DOMAIN–CONTAINING RECEPTORS – DECISIONS BETWEEN SUICIDE AND FIRE |
31 |
TNF-R1 signaling complex
|
RIP1 |
TRADD |
|
|
TRAF2/5 |
TAB2 |
TAK1 |
|
TAB1 |
|
|
|
|
|
NEMO IKKβ |
|
cIAP1/2 |
|
|
|
IKKα |
|
|
NF-κB JNK p38
Gene induc on
Figure 3-3. Schematic representation of immunostimulatory, proinflammatory signaling by the TNF-R and DR3 systems. Binding of TNF and TL1A to their respective receptors leads to receptor trimerization and formation of a receptor signaling complex. First the adaptor protein TNF-R1–associated death domain (TRADD) is recruited via its DD to the DD of the receptor. TRADD then serves as an assembly platform for binding of TRAF2, cIAP1/2, and the receptor interacting kinase 1 (RIP1). TRAFs and cIAPs conjugate ubiquitin chains to various proteins in the complex, which allows for the recruitment of further signaling proteins, including the TAK/TAB and the IKK complexes, ultimately leading to activation of NF-κ B and the JNK and p38 MAP kinase pathways. See Color Plate 5.
efficient TNF-induced NF-κB activation because NF-κB essential modulator (NEMO) binds strongly to linear but only weakly to K63-linked ubiquitin chains. Together, K63-linked and linear polyubiquitination of different components of the TNF-RSC result in stable TNF-RSC formation and thereby enable the events that ultimately lead to activation of NF-κB and the JNK and p38 MAP kinase pathways. The molecular processes that lead to the activation of these pathways have been extensively reviewed in the past (Hayden and Ghosh, 2008; Wajant et al., 2003). The discovery of LUBAC as a novel integral component of the TNF-RSC and linear ubiquitination as a central player in the organization of this protein complex will undoubtedly substantially affect our current view of how these processes are regulated. It will be exciting to unravel these mechanisms at the molecular level and discover how they control the function of TNF.
3.1.2. TNF and TNF blockers in the clinic
Soon after the isolation of TNF, it became clear that the systemic administration of TNF is highly toxic as
a result of the extreme cytokine production it induces. This inflammation-like syndrome prevented the further development of TNF for systemic use. However, a technique called isolated limb perfusion (ILP) has been developed by Ferdinand Lejeune in Lausanne, Switzerland (Lejeune et al., 1995). ILP facilitates local and exclusive administration of TNF to, for example, an arm or leg of a patient with cancer. ILP with TNF in combination with chemotherapeutic drugs led to complete response rates in some patients with sarcomas and melanomas on extremities and showed improved penetrance of the cytostatic drugs melphalan and doxorubicin into tumors in animal models. The possibility to inhibit any leaked TNF in the rest of the body with therapeutic TNF blockers, which are now available, may help to overcome the limitations posed so far on ILP by the detrimental effects of potential TNF leakage. Interestingly, TNF specifically disrupts tumor-supporting blood vessels while sparing normal blood vessels. It would be interesting to examine whether endothelial cells in the tumor-associated neovasculature are particularly sensitive to TNF-induced apoptosis and what the biochemical
32 |
|
|
HENNING WALCZAK AND CHAHRAZADE KANTARI |
|
Primarily apopto c signaling systems |
Primarily immunos mulatory, proinflammatory |
|||
(CD95 and TRAIL systems) |
signaling systems (TNF and DR3 systems) |
|||
Complex I |
|
|
Complex I |
|
FADD |
RIP1 |
TRADD |
|
|
|
|
|||
Complex II |
|
|
TRAF2/5 |
Complex II |
Caspase- 8 |
TAB2 |
TAK1 |
|
|
|
TAB1 |
|
|
|
NEMO |
NEMOIKKβ |
|
cIAP1/2 |
|
|
IKKα |
|
|
|
NF-κB
MAPK
APOPTOSIS |
Gene induc on |
APOPTOSIS |
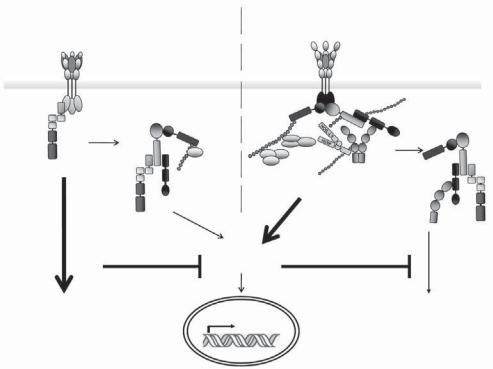
Figure 3-4. Complex I and complex II: spatial dissociation between proapoptotic and proinflammatory signaling in death receptor signal transduction. For both the CD95 and TRAIL systems, as well as the TNF and DR3 systems, the complex defined as complex I is the protein complex that forms at the plasma membrane and exerts the primary function of the respective receptor (i.e., apoptosis for CD95 and TRAIL-R1/R2 and gene induction via NF-κB and MAPK activation by TNFR-R1 and DR3). By an undetermined mechanism, the primary adaptor protein for the di erent complexes I dissociates from the DD of the respective receptor, together with a number of other proteins assembled in complex I (but without the receptor), and recruits additional proteins from the cytosol. This complex II then triggers the secondary function of each receptor. In the case of proapoptotic receptors, this is gene induction via activation of NF-κB and the MAP kinases pathways; in the case of the primarily immunostimulatory, proinflammatory receptors, it is induction of apoptosis. Successful completion of the respective primary signal interferes with the execution of the respective secondary signal. See Color Plate 6.
basis for this is. ILP is approved for unresectable soft tissue sarcoma, and it has also been successfully applied in the treatment of various other local tumors. The success of this technique proved that TNF can be used to treat cancer, albeit only when administration is locally restricted and by exerting its killing activity on an unexpected cellular target. Hence successful TNF treatment has mainly become a matter of targeted delivery to the tumor site.
A molecular way of achieving targeted delivery is to create fusion proteins in which TNF is conjugated to antibody fragments or natural ligands that specifically recognize surface proteins on tumor cells or in the tumor stroma. Using this technique, significant killing of tumor cells has been obtained, and exciting new recombinant proteins are currently being investigated. As an example, a melanoma-specific antibody conjugated to recombinant human TNF exhibited very good killing activity against TNF-resistant melanoma cells, both in
vitro and in vivo. Thus these types of fusion proteins or conjugates may not only result in more effective tumor targeting, but may also enhance the killing activity of TNF, most likely by providing membrane fixation and thereby enabling higher order receptor cross-linking on the target cell. Similar fusion proteins have also been constructed with CD95L and TRAIL. The preclinical results obtained with some of the proteins are very encouraging.
The by far most important clinical development in the TNF field to date emerged from the initially discouraging observation that TNF exerts an inflammatory response. When scientists started investigating the upside of this, they realized that interference with this response can be used to treat inflammatory conditions. In the beginning it was thought that sepsis could be targeted by inhibiting TNF. However, it was overlooked that Daniela Mannel¨ and her team, then in Heidelberg, Germany, had already shown that TNF plays an ambiguous
