Neutron Scattering in Biology - Fitter Gutberlet and Katsaras
.pdf21 Internal Dynamics of Proteins and DNA |
499 |
is usually beyond the resources of most laboratories. However, the European Community has recently established a laboratory for large-scale production of deuterated molecules [72]. These problems with sample size should also be alleviated when new spallation neutron sources become available. It is expected that they will provide signals 100 (in the most optimistic estimate even 1,000) times stronger than current instruments. With these sources samples required for scattering measurements might be reduced to 1–10 mg, i.e., to the amount available to many biochemical laboratories.
Precise selective deuteration of biological macromolecules will open tremendous possibilities for analysis of motion of active sites, relative motions of secondary structures, for studies of dynamic heterogeneity in proteins (see, for example, ideas formulated in [73, 74]) and many other questions. Completely deuterated proteins will open the possibility for analysis of coherent scattering that will allow the analysis of pair-correlation functions.
We also expect that significant advantages in understanding protein and DNA dynamics might be achieved by using ideas developed in the field of dynamics of complex systems. Significant progress in this field has been achieved during the last two decades. It was based on developments of two approaches
– the energy landscape and mode coupling theory. In particular, it emphasizes that the dynamics of liquid systems can be separated into three regimes [65]:
(i) simple liquid behavior at very high temperatures; (ii) onset of some collective (or coupled) dynamics, which happens below some onset temperature T . This region is usually called “energy landscape influenced” and extends down to the dynamic crossover temperature TC; (iii) crossover from liquid-like to solid-like dynamics on the molecular scale which occurs at TC; the liquid becomes nonergodic on a short time scale. The region at T < TC is called “energy landscape dominated”. The first regime has no interest for biological macromolecules because of its high temperature. In our view, biomolecules in their native environment are in the second regime. In this regime, relaxation over large barriers does not dominate the dynamics. The hierarchy of conformational substates available to the native protein gives rise to a rough energy landscape with a broad distribution of barrier heights. The current view is that the system can relax or undergo conformational changes by negotiating its way around the high-energy barriers in the multidimensional energy landscape. It does not have to jump over these barriers. Below TD, however, the system becomes trapped in deep energy minima of the landscape. As a result, harmonic vibrations dominate S(Q, ν) and over-barrier relaxation becomes the dominant mechanism of conformational motion in the third regime. Application of these ideas to analysis of dynamics of proteins and DNA might be very helpful in understanding microscopic mechanisms of molecular motions. Neutron spectroscopy continues to play a key role in the analysis of dynamics of complex systems and with the developments in new neutron sources and the production of deuterated proteins should receive increased attention from the biophysical and biochemical community in future.
500 A.P. Sokolov et al.
Acknowledgements
AS is grateful to NSF (DMR-0315388) for financial support, and to ILL and NIST Neutron Center for assistance with neutron measurements.
References
1.J. Perez, J.M. Zanotti, D. Durand, Biophys. J. 77 (1999), 454
2.M. Tarek, D.A. Neumann, D.J. Tobias, Chem. Phys. 292 (2003), 435
3.A.P. Sokolov, H. Grimm, A. Kisliuk, A.J. Dianoux, J. Biol. Phys. 27 (2001), 313
4.J.H. Roh, G. Caliskan, A.P. Sokolov, R. Gregory, et al. (unpublished)
5.C.L. Brooks III, M. Karplus, B.M. Pettitt, in Proteins: A Theoretical Perspective of Dynamics, Structure, and Thermodynamics (Wiley Interscience, 1988) p. 19
6.S. Longeville, W. Doster, G. Kali, Chem. Phys. 292 (2003), 413
7.W. Doster, S. Cusack, W. Petry, Phys. Rev. Lett. 65 (1990), 1080
8.J. Fitter, R.E. Lechner, N.A. Dencher, Biophys. J. 73 (1997), 2126
9.A.P. Sokolov, H. Grimm, R. Kahn, J. Chem. Phys. 110 (1999), 7053
10.M. Tarek, D.J. Tobias, J. Chem. Phys. 115 (2001), 1607
11.F.G. Parak, Rep. Prog. Phys. 66 (2003), 103
12.G. Caliskan, A. Kisliuk, A. Sokolov, J. Non-Crystallogr. Sol. 307–310 (2002), 868
13.C.P. Lindsey, G.D. Patterson, J. Chem. Phys. 73 (1980), 3348
14.J.L. Green, J. Fan, C.A. Angell, J. Am. Chem. Soc. 98 (1994), 13780
15.C.A. Angell, Science 267 (1995), 1924
16.W. Doster, M. Settles, in Workshop on Hydration Processes in Biology: Theoretical and Experimental Approaches, M.C. Bellissent-Funel (Ed.) (Les Houches, IOS Press, 1999) p. 177
17.D. Richter, et al., J. Phys. Condens. Matter 11 (1999), A297
18.J. Colmenero, et al., J. Phys. Condens. Matter 11 (1999), A363
19.F. Volino, A.J. Dianoux, Mol. Phys. 41 (1980), 271
20.A.L. Tournier, J.C. Smith, Phys. Rev. Lett. 91 (2003), 208106
21.J.A. Rupley , P. Yang, G. Tollin, in Water in Polymers, S.P. Rowland (Ed.) (Am. Chem. Soc., Washington, DC, 1980) pp. 111–132
22.O.V. Belonogova, E.N. Frolov, S.A. Krasnopol’skaya, B.P. Atanasov, V.K. Gins, E.N. Mukhin, A.A. Levina, A.P. Andreeva, G.I. Likhtenshtein, V.I. Goldanskii, Dokl. Akad. Nauk USSR 241 (1978), 219
23.G.I. Likhtenshtein, V.R. Bogatyrenko, A.V. Kulikov, Appl. Magn. Reson. 4 (1993), 513
24.F. Parak, Methods Enzymol. 127 (1986), 196
25.G.B. Strambini, E. Gabellieri, Photochem. Photobiol. 39 (1984), 725
26.N.K. Shah, R.D. Ludescher, Photochem. Photobiol. 58 (1993), 169
27.R.B. Gregory, M. Gangoda, R.K. Gilpin, W. Su, Biopolymers 33 (1993), 513
28.P.L. Poole, J.L. Finney, Int. J. Biol. Macromol. 5 (1983), 308
29.J.E. Schinkel, N.W. Downer, J.A. Rupley, Biochemistry 24 (1985), 352
30.S. Bone, R.J. Pethig, Mol. Biol. 181 (1985), 323
21 Internal Dynamics of Proteins and DNA |
501 |
31.R. Pethig, in Protein–Solvent Interactions, R.B. Gregory (Ed.) (Marcel Dekker, New York, 1995) pp. 265–288
32.V.I. Goldanskii, Y.F. Krupyanskii: Rev. Biophys. 22 (1989), 39; V.I. Goldanskii, Y.F. Krupyanskii, in Protein-Solvent Interactions, R.B. Gregory (Ed.) (Marcel Dekker, New York, 1995) pp. 289–326
33.M. Diehl, W. Doster, W. Petry, A. Schulte, Biophys. J. 73 (1997), 2726
34.R.B. Gregory, in Protein–Solvent Interactions, R.B. Gregory (Ed.) (Marcel Dekker, New York, 1995) pp. 191-264
35.R.W. Lumry, A. Rosenberg, Colloques. Int. CNRS 246 (1975), 53
36.M. Tarek, D.J. Tobias, Phys. Rev. Lett. 88 (2002), 138101
37.A.L. Tournier, J. Xu, J.C. Smith, Biophys. J. 85 (2003), 1871
38.D. Vitkup, D. Ringe, G.A. Petsko, M. Karplus, Nat. Struct. Biol. 7 (2000), 34–38
39.A. Ansari, C.M. Jones, E.R. Henry, J. Hofrichter, W.A. Eaton, Science 256 (1992), 1796
40.T. Kleinert, et al., Biochemistry 37 (1998), 717
41.H. Lichtenegger, et al., Biophys. J. 76 (1999), 414
42.B. Gavish, M.M. Werber, Biochem. 18 (1979), 1269
43.D. Beece, L. Eisenstein, H. Frauenfelder, D. Good, M.C. Marden, L. Reinish, A.H. Reynolds, L.B. Sorensen, K.T. Yue, Biochem. 19 (1980), 5147
44.M. Settles, W. Doster, F. Kremer, F. Post, W. Schirmacher, Philos. Mag. 65 (1992), 861
45.R.H. Austin, K.W. Beeson, L. Eisenstein, H. Frauenfelder, I.C. Gunsalus, Biochem. 14 (1975), 5355
46.B. Gavish, S. Yedgar, in Protein–Solvent Interactions, R.B. Gregory (Ed.) (Marcel Dekker, New York, 1995) pp. 343–373
47.S.N. Timashe , in Protein–Solvent Interactions, R.B. Gregory (Ed.) (Marcel Dekker, New York, 1995) pp. 445–482
48.R. Gregory, Biopolymers 27 (1988), 1699
49.W. Doster, S. Cusak, W. Petry, Nature 337 (1989), 754
50.B.F. Rasmussen, et al., Nature 357 (1992), 423
51.A.M. Tsai, D.A. Neumann, L.N. Bell, Biophys. J. 79 (2000), 2728
52.M. Ferrand, A.J. Dianoux, W. Petry, G. Zaccai, Proc. Natl Acad. Sci. USA 90 (1993), 9668
53.R.M. Daniel, J.C. Smith, M. Ferrand, S. Hery, R. Dunn, J. Finney, Biophys. J. 75 (1998), 2504
54.L. Cordone, M. Ferrand, E. Vitrano, G. Zaccai, Biophys. J. 76 (1999), 1043
55.W. Gotze, L. Sjogren, Rep. Prog. Phys. 55 (1992), 241
56.see, for example, Transp. Theory Stat. Phys. 24 (1995), Special issue devoted to Relaxation Kinetics in Supercooled Liquids – Mode Coupling Theory and Its Experimental Tests
57.W. Stefen, A. Patkowski, H. Glaser, G. Meier, E.W. Fischer, Phys. Rev. E 49 (1994), 2992
58.A.P. Sokolov, W. Ste en, E. R¨ossler, Phys. Rev. E 52 (1995), 5105
59.A.P. Sokolov, Science 273 (1996), 1675; Endeavour 21 (1997), 109
60.A.P. Sokolov, J. Hurst, D. Quitmann, Phys. Rev. B 51 (1995), 12865
61.E. R¨ossler, A.P. Sokolov, A. Kisliuk, D. Quitmann, Phys. Rev. B 49 (1994), 14967
62.A.P. Sokolov, J. Non-Crystallogr. Solids 235–237 (1998), 190
502A.P. Sokolov et al.
63.N.V. Surovtsev, J. Wiedersich, V.N. Novikov, E. R¨ossler, A.P. Sokolov, Phys. Rev. B 58 (1998), 14888
64.J. Fitter, Biophys. J. 76 (1999), 1034
65.P.G. Debenedetti, F.H. Stillinger, Nature 410 (2001), 259
66.J.W. Mack, M.G. Usha, J. Long, R.G. Gri n, R.J. Wittebort, Biopolymers 53 (2000), 9
67.G. Caliskan, A. Kisliuk, V.N. Novikov, A.P. Sokolov, J. Chem. Phys. 114 (2001), 10189
68.G. Caliskan, A. Kisliuk, A. Tsai, C. Soles, A.P. Sokolov, J. Chem. Phys. 118 (2003), 4230
69.M.T. Cicerone, A. Tellington, L. Trost, A. Sokolov, Bioprocess Int. 1 (2003), 2
70.V. Reat, H. Patzelt, C. Pfister, M. Ferrand, D. Oesterhelt, G. Zaccai, PNAS USA 95 (1998), 4970
71.A. Orecchini, A. Paciaroni, A.R. Bizzarri, S. Canistraro, J. Phys. Chem. B 106 (2002), 7348
72.Information about the ILL-EMBL Deuteration Laboratory can be found on the web: www.ill.fr/deuteration
73.Y. Zhou, D. Vitkup, M. Karplus, J. Mol. Biol. 285 (1999), 1371
74.S. Dellerue, A.J. Petrescu, J.C. Smith, M.C. Bellissent-Funel, Biophys. J. 81 (2001), 1666

22
Structure and Dynamics
of Model Membrane Systems Probed
by Elastic and Inelastic Neutron Scattering
T. Salditt, M.C. Rheinst¨adter
22.1 Introduction
Phospholipid membranes are intensively studied as simple model systems to understand fundamental structural and physical aspects of their much more complex biological counterparts [1]. The lateral structure of membranes, including both height and compositional fluctuations, remains an important experimental challenge of present-day biophysics, concerning in particular the biologically relevant fluid Lα state, where the material softness compromises the use of scanning probe microscopy. Neutron scattering can contribute to the elucidation of the molecular structure, as is well documented in the literature (see, e.g., [2]).
Dynamical properties are often less well understood in biomolecular systems, but are important for many fundamental biomaterial properties, e.g., elasticity properties and interaction forces. Furthermore, lipid membrane dynamics on small molecular length scales determines or strongly a ects functional aspects, like di usion and parallel or perpendicular transport through a bilayer. The specific advantages of neutron scattering to study fluctuations of phospholipid membranes on lateral length scales between several micrometer down to a few ˚Angstr¨oms can give unique insights.
The present chapter concentrates on mainly two neutron scattering techniques, which give information on very di erent types of dynamics: (i) nonspecular neutron reflectivity (NSNR) as a tool to probe thermal fluctuations of lipid bilayers on mesoscopic length scales, and (ii) inelastic neutron scattering (INS) for studies of the short-range collective motions in the acyl chains. The methods can also be applied to more complex model systems, including lipid – peptide and lipid–protein mixtures, as well as in some cases to real biological membranes like purple membranes.
The chapter is so organized that at first sample preparation and sample environment for multilamellar lipid phases on solid support is presented. Next, specular neutron reflectivity (SNR) and NSNR from lipid membranes is described. Mainly work on pure lipid phases is presented and reviewed, a
504 T. Salditt, M.C. Rheinst¨adter
few examples relate to structure and interaction of the antimicrobial peptide Magainin 2 in phosphocholine bilayers, as an example of how the methods presented can be extended and applied to probe lipid – peptide interaction or more generally the interaction of bilayers with membrane-active molecules (such as stereols, peptides, and proteins). Afterward recent advances are highlighted in the application of classical INS at triple-axis spectrometers (TAS) to study the short-range dynamics of lipid bilayers, e.g., the collective motion of the acyl chains. The chapter closes with a summary and conclusions.
22.2 Sample Preparation and Sample Environment
Highly oriented multilamellar bilayers for neutron reflectivity can be easily prepared on cleaned silicon wafers ((1 1 1)-orientation or (1 0 0)-orientation) by spreading from organic solution [3]. The wafers are cleaned by subsequent washing in methanol and ultrapure water (specific resistivity ≥ 18 MΩ cm), and made hydrophilic by washing in a 5-molar solution of KOH in ethanol for about a minute, or alternatively by plasma etching. Lipid and/or peptide components are codissolved in the desired ratio (molar ratio P/L) in trifluoroethanol (TFE) or (1:1) TFE–chloroform mixtures at concentrations between 10 and 40 mg ml−1, depending on the total mass to be deposited.
After a slow evaporation process avoiding film rupture, remaining traces of solvent in the sample are removed by exposing the samples to high vacuum overnight. Then the films are rehydrated in a hydration chamber and tempered above the main phase transition. The orientational alignment of such multilamellar stack with respect to the substrate (mosaicity) is typically better than 0.01◦. A very low mosaicity is a prerequisite to apply interfacesensitive scattering techniques. The lateral domains sizes are in the range of 100 m, exhibiting a broad distribution in the total number N of the bilayers.
For measurements on samples immersed in water thick (1 cm), polished silicon blocks are used, where the neutron beam can be coupled into the substrate without refraction from the side. The beam then impinges at grazing incidence onto the sample with the incoming and reflected beam path in silicon rather than in water, avoiding incoherent scattering in H2O or D2O/H2O mixtures (Fig. 22.1b). The low mosaicity of the samples prepared was preserved after immersing the samples in water.
For INS experiments highly oriented membrane stacks were prepared essentially in the same way as presented earlier [4]. However, ten or more such wafers separated by small air gaps are combined and aligned with respect to each other to create a “sandwich sample” consisting of several thousands of highly oriented lipid bilayers (total mosaicity of about 0.6◦), with a total mass of several hundred milligrams of deuterated phospholipid, to maximize the scattering volume. Figure 22.1c shows a photograph of a sample for the inelastic neutron experiments.
During the neutron experiments, the solid-supported multilamellar films are kept in temperature and hydration controlled chambers. For measurements

22 Structure and Dynamics of Model Membrane Systems |
505 |
(a) |
(b) |
|
Si |
(c) |
(d) |
x N
Si
Fig. 22.1. (a) Schematic of the chamber used for the neutron experiments to control temperature and humidity of the bilayers. (b) Samples immersed in water were applied on 1 cm thick Si wafers. The neutron beam was then coupled into the wafer from the side. (c) Photograph of the “sandwich sample” prepared for the inelastic neutron experiments. (d) Hydrated multilamellar sample with layers of water (H2O/D2O) between the bilayers, respectively
carried out at partial hydration with the bilayers facing D2O vapor, a chamber with two concentric high-purity aluminum cylinders was used, see Fig. 22.1a. The inner cylinder was heated or cooled by a flow of oil, connected to a temperature-controlled reservoir. The space between the two cylinders was evacuated to minimize heat conduction. The temperature was measured close to the sample holder by a Pt100 sensor, indicating a thermal stability of better than 0.02 K over several hours. At the bottom of the inner cylinder, a water reservoir was filled with salt-free Millipore water, such that the sample was e ectively facing a vapor phase of nominally 100% relative humidity.
Despite the nominally full hydration condition, bilayer samples of DMPC were typically swollen only up to a repeat distance of only d 50–55 ˚A in the fluid Lα-phase, i.e., were only partially hydrated. This limited swelling of solid-supported lipid films is well known as the so-called vapor-pressure paradox, and has recently been explained by Katsaras and coworkers on the basis of small temperature gradients in the chamber [5,6]. Accordingly, it was demonstrated that chambers of suitable design do not show this e ect, and that bilayers can be swollen to equilibrium (full hydration) from the vapor phase.
Alternatively, a chamber operating at full hydration has been used, where the bilayers are immersed in water, and the beam impinges through a thick
22 Structure and Dynamics of Model Membrane Systems |
507 |
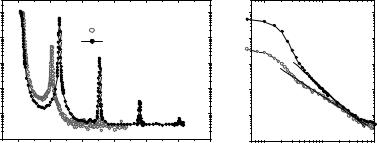
the second-order Bragg peak. At the same time the increase in thermal fluctuations at full hydration leads to a significant damping of higher-order peak, so that only one (strong) Bragg peak is observed. At partial hydration more than four Bragg peaks can be measured, depending on the exact relative humidity (osmotic pressure).
The correct analysis of X-ray and neutron reflectivity relies on very low mosaicity (narrow orientational distribution of domains). A necessary condition is a clear distinction between specular and nonspecular scattering components. The data analysis and modeling of the measured reflectivity should be based on an appropriate scattering theory, such as the fully dynamical Parratt algorithm (taking into account multiple reflections) or on the semikinematical reflectivity pioneered by Als-Nielsen [12]. The observation of a region of total external reflection and hence of the critical angle αc allows for the determination of the scattering length density profile on an absolute scale. Moreover, since the full qz -range can be used for data analysis by fitting the reflectivity curve to a parametrized model of the density profile [13], a reasonable resolution in ρ(z) can also be reached for fully hydrated systems. Furthermore, the phase problem is reduced, since the change of sign in the bilayer form factor (real valued due to centro-symmetry) is often accompanied by an observable cusp in the (continuously measured) reflectivity curve. Alternatively, phasing can be performed by the so-called swelling method. The advantage of full qz -fitting has also been demonstrated in bulk (SAXS) studies, see for example [14].
In most published studies of oriented bilayers, however, only the integrated Bragg peaks of the multilamellar samples are used for data analysis, and the one-dimensional density profile ρ(z) is computed by Fourier synthesis using a discrete set of Fourier coe cients fn as described in [15, 16]. In this approach, the exact relation between Bragg peak intensity and the Fourier co- e cients fn is an open problem for which there may not even exist a general solution. Taking into account e ects of absorption, polarization, specular and nonspecular Fresnel reflectivity components, illumination correction, etc., a widely used correction factor is In = |fn|2/qz , where qz−1 is termed a Lorentz factor for oriented bilayers. In the absence of a rigorous theoretic derivation, such a correction is at best empirical. Furthermore, the two scattering contributions of specular and nonspecular scattering are often measured in the same scan, adding up the background and making quantitative analysis questionable, since both contributions are governed by a di erent qz dependence. However, the peak-to-peak distance of a reconstructed bilayer profile is luckily relatively stable against variations in data analysis like di erent choices of Lorentz factors.
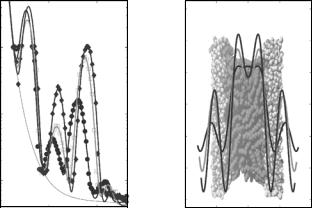
Figure 22.3a shows DMPC reflectivity curves measured at a rather low resolution on a TAS, along with the results for the density profiles obtained by a Fourier synthesis approach (Fig. 22.3b). The data have been recorded as function of T below and above the main phase transition, simultaneously with inelastic data discussed in Sect. 22.5. The temperature-dependent structural