
- •Contents
- •Contributors
- •Preface
- •1 Introduction, with the biological basis for cell mechanics
- •Introduction
- •The role of cell mechanics in biological function
- •Maintenance of cell shape
- •Cell migration
- •Mechanosensing
- •Stress responses and the role of mechanical forces in disease
- •Active cell contraction
- •Structural anatomy of a cell
- •The extracellular matrix and its attachment to cells
- •Transmission of force to the cytoskeleton and the role of the lipid bilayer
- •Intracellular structures
- •Overview
- •References
- •2 Experimental measurements of intracellular mechanics
- •Introduction
- •Forces to which cells are exposed in a biological context
- •Methods to measure intracellular rheology by macrorheology, diffusion, and sedimentation
- •Whole cell aggregates
- •Sedimentation of particles
- •Diffusion
- •Mechanical indentation of the cell surface
- •Glass microneedles
- •Cell poker
- •Atomic force microscopy
- •Mechanical tension applied to the cell membrane
- •Shearing and compression between microplates
- •Optical traps
- •Magnetic methods
- •Twisting of magnetized particles on the cell surface and interior
- •Passive microrheology
- •Optically detected individual probes
- •One-particle method
- •Two-particle methods
- •Dynamic light scattering and diffusing wave spectroscopy
- •Fluorescence correlation spectroscopy
- •Optical stretcher
- •Acoustic microscopy
- •Outstanding issues and future directions
- •References
- •3 The cytoskeleton as a soft glassy material
- •Introduction
- •Magnetic Twisting Cytometry (MTC)
- •Measurements of cell mechanics
- •The structural damping equation
- •Reduction of variables
- •Universality
- •Scaling the data
- •Collapse onto master curves
- •Theory of soft glassy rheology
- •What are soft glassy materials
- •Sollich’s theory of SGMs
- •Soft glassy rheology and structural damping
- •Open questions
- •Biological insights from SGR theory
- •Malleability of airway smooth muscle
- •Conclusion
- •References
- •4 Continuum elastic or viscoelastic models for the cell
- •Introduction
- •Purpose of continuum models
- •Principles of continuum models
- •Boundary conditions
- •Mechanical and material characteristics
- •Example of studied cell types
- •Blood cells: leukocytes and erythrocytes
- •Limitations of continuum model
- •Conclusion
- •References
- •5 Multiphasic models of cell mechanics
- •Introduction
- •Biphasic poroviscoelastic models of cell mechanics
- •Analysis of cell mechanical tests
- •Micropipette aspiration
- •Cells
- •Biphasic properties of the pericellular matrix
- •Indentation studies of cell multiphasic properties
- •Analysis of cell–matrix interactions using multiphasic models
- •Summary
- •References
- •6 Models of cytoskeletal mechanics based on tensegrity
- •Introduction
- •The cellular tensegrity model
- •The cellular tensegrity model
- •Do living cells behave as predicted by the tensegrity model?
- •Circumstantial evidence
- •Prestress-induced stiffening
- •Action at a distance
- •Do microtubules carry compression?
- •Summary
- •Examples of mathematical models of the cytoskeleton based on tensegrity
- •The cortical membrane model
- •Tensed cable nets
- •Cable-and-strut model
- •Summary
- •Tensegrity and cellular dynamics
- •Conclusion
- •Acknowledgement
- •References
- •7 Cells, gels, and mechanics
- •Introduction
- •Problems with the aqueous-solution-based paradigm
- •Cells as gels
- •Cell dynamics
- •Gels and motion
- •Secretion
- •Muscle contraction
- •Conclusion
- •Acknowledgement
- •References
- •8 Polymer-based models of cytoskeletal networks
- •Introduction
- •The worm-like chain model
- •Force-extension of single chains
- •Dynamics of single chains
- •Network elasticity
- •Nonlinear response
- •Discussion
- •References
- •9 Cell dynamics and the actin cytoskeleton
- •Introduction: The role of actin in the cell
- •Interaction of the cell cytoskeleton with the outside environment
- •The role of cytoskeletal structure
- •Actin mechanics
- •Actin dynamics
- •The emergence of actin dynamics
- •The intrinsic dynamics of actin
- •Regulation of dynamics by actin-binding proteins
- •Capping protein: ‘decommissioning’ the old
- •Gelsolin: rapid remodeling in one or two steps
- •β4-thymosin: accounting (sometimes) for the other half
- •Dynamic actin in crawling cells
- •Actin in the leading edge
- •Monomer recycling: the other ‘actin dynamics’
- •The biophysics of actin-based pushing
- •Conclusion
- •Acknowledgements
- •References
- •10 Active cellular protrusion: continuum theories and models
- •Cellular protrusion: the standard cartoon
- •The RIF formalism
- •Mass conservation
- •Momentum conservation
- •Boundary conditions
- •Cytoskeletal theories of cellular protrusion
- •Network–membrane interactions
- •Network dynamics near the membrane
- •Special cases of network–membrane interaction: polymerization force, brownian and motor ratchets
- •Network–network interactions
- •Network dynamics with swelling
- •Other theories of protrusion
- •Numerical implementation of the RIF formalism
- •An example of cellular protrusion
- •Protrusion driven by membrane–cytoskeleton repulsion
- •Protrusion driven by cytoskeletal swelling
- •Discussion
- •Conclusions
- •References
- •11 Summary
- •References
- •Index
188J.L. McGrath and C.F. Dewey, Jr.
Ca2+ requirements, it is unclear if these steady-state effects are mediated by intermittent and localized gelsolin activity, or constant but low levels of activity. Gelsolin also has a fascinating role in apoptosis where caspases cleave the protein to create an unregulated severing peptide (Kothakota et al., 1997). Continuous severing by this peptide helps create a mechanically compromised cell that eventually detaches from its substrate (Kothakota et al., 1997).
β4-thymosin: accounting (sometimes) for the other half
The 15 kD protein β4-thymosin is present in mammalian cells at levels that equal or exceed actin itself (Safer and Nachmias, 1994). Its discovery appeared to resolve the critical question of how mammalian cells maintained nearly half of their actin in an unpolymerized form despite intracellular conditions that favored polymerization. β4-thymosins are unstructured in solution and partially wrap around the G-actin monomer to control its associations (Safer et al., 1997). β4-thymosins bind to ATP- G-actin (but not ADP-G-actin) with an affinity comparable to the pointed end ATP critical concentration, but less than the barbed end ATP critical concentration (Carlier et al., 1993). In this way when barbed ends are mostly capped, β4-thymosin functions as a ‘sequestering’ protein that maintains a pool of nonfilamentous actin, but when free barbed ends are abundant the pool diminishes. Profilin can compete for the ATP-G-actin pool maintained by β4-thymosin (Carlier et al., 1993), possibly by emerging with the charged monomer after forming a complex that includes all three proteins (Yarmola et al., 2001). Thus with or without profilin, β4-thymosin helps to reserve ATP-G-actin for future assembly at barbed ends. The high concentration of β4-thymosin can support extensive and sudden conversions from G-actin to filaments. This conversion is likely occurring in the dramatic polymerization-induced shape change of both platelets (Safer et al., 1990) and neutrophils (Cassimeris et al., 1992), both of which contain abundant β4-thymosin/G-actin complex at rest. However β4-thymosin apparently is not required for the continuous shape change during crawling, because motile amoebae are thought to be void of thymosin-family proteins (Pollard et al., 2000).
Dynamic actin in crawling cells
In this section we explore a most conspicuous and well-studied function of the actin cytoskeleton: its ability to serve as the engine for cell crawling. By driving the expansion of the plasma membrane in the direction of cell advancement, actin polymerization initiates the crawling cycle (Fig. 9-8). The networks formed by polymerization evolve to structures that provide mechanical support for cell extensions; that link the cell to its substrate; and that support the myosin-based contractions needed for cell translation. The network must also disassemble to recycle its constituents for further rounds of assembly. Thus the actin network at the leading edge of motile cells provides both the structure and the forces needed for crawling (see Fig. 9-9). Here we review the current understanding of the geometry and dynamics of these networks, and address the important question of how polymerization might lead to pushing forces.
Cell dynamics and the actin cytoskeleton |
189 |
||
|
Cell body |
Leading edge |
|
|
|
|
|
Tail |
|
protrusion |
|
Adhesion
site
1)
Extracellular
matrix
2)
3)
4)
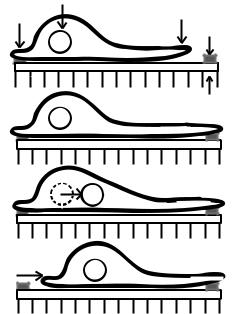
Fig. 9-8. The four steps in cell migration. The classic schematic of crawling breaks the process down into a four-step cycle. The cycle begins with the protrusion of the leading edge driven by actin polymerization. The extended cell forms new attachments in advance of its body and then contracts against this attachment to break tail adhesions and translate forward. From Mitchison and Cramer, 1996.
Actin in the leading edge
The extension of the plasma membrane that interrogates new regions of substrate can come in several forms. Mammalian cells crawling in culture environments extend both finger-like projections, called fillopodia, and broad, thin, veil-like projections called lamellipodia. Which structure occurs more frequently is a strong function of cell type and substrate conditions (Pelham and Wang, 1997). Cells that crawl in amoeboid fashion – a class that includes the leukocytes of the mammalian immune system – use bulkier protrusions known as pseudopodia. By all accounts, the initiator of filament assembly in each of these cellular protrusions is the Arp2/3 complex activated by membrane-bound WASp/Scar family proteins. Pollard and Borisy have offered the most detailed proposal for how actin networks evolve in lamellipodia (Pollard and Borisy, 2003), and filopodia appear to be triggered from rearrangements of a lamellipodial network (Svitkina et al., 2003). Because the three-dimensional character of pseudopodia makes them less amenable to ultrastructural and fluorescence studies, far less is known about the geometry and dynamics of pseudopodial networks.
Synthesizing data from electron micrographs of cytoskeletal structure in the lamellipodia of fast-moving keratocytes (Svitkina and Borisy, 1999), immunochemical analysis of Arp2/3 complex, ADF/cofilin, and capping protein location in these same samples (Svitkina and Borisy, 1999), and live cell fluorescence data revealing regions of actin assembly and disassembly in fibroblasts (Watanabe and Mitchison, 2002),
190 J.L. McGrath and C.F. Dewey, Jr.
A
B
G-actin
F-actin
polymerization rate |
(monomers / s) |
|
monomer |
concentration |
( µM) |
filament velocity |
relative toleading edge |
( µm / s) |
diffusive transport
retrograde flow
100,000
0 -3,000
30
15
0
0 -0.025
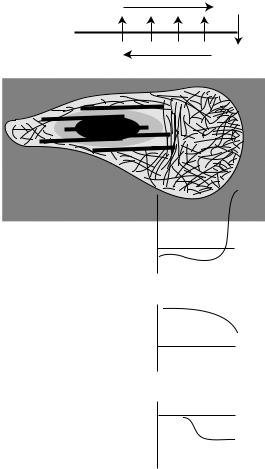
Fig. 9-9. Actin dynamics in crawling cells. (A) For steady crawling, actin polymerization and depolymerization must complete a balanced cycle. However, the demand for assembling monomers at the leading edge causes a spatial segregation of these processes and flows. Actin that assembles at the leading edge but does not incorporate into a protrusion flows in retrograde fashion as it is disassembled. The emerging G-actin population is presumably returned to the leading edge by diffusion. (B) The graphs show possible profiles for polymerization, monomer concentration, and retrograde flow across the front of a crawling cell. The calculated numbers combine measurements of actin dynamics in fibroblasts (Vallotton et al., 2004; Watanabe and Mitchison, 2002) and the leading edge monomer demand for these cells (Abraham et al., 1999), and assumes monomer is returned by diffusion with a diffusivity of 6 × 10−8 cm2/s. From McGrath et al., 1998a.
Pollard and Borisy propose the lamellipodia are filled with a neatly segregated, highly dynamic network (Pollard and Borisy, 2003). In their model, filaments are first generated by activated Arp2/3 complex to form brush-like networks within one micron of the advancing plasma membrane. The newly born filaments remain short because their growth is rapidly terminated by abundant capping protein. The continuous assembly of ATP-actin at the leading edge explains why the ADP-actin specific ADF/cofilins are excluded from this region (Svitkina and Borisy, 1999). ADF/cofilin-family proteins
Cell dynamics and the actin cytoskeleton |
191 |
take up residence at distances one micron and more from the leading edge where they bind to aged ADP-actin, disassembling the network into monomers for rapid recycling and further polymerization (Svitkina and Borisy, 1999). More recent and sophisticated analysis of actin filament dynamics are consistent with a segregation of the lamellipod into a 1 micron membrane-proximal region dominated by polymerization and an immediately adjacent region where significant depolymerization occurs (Vallotton et al., 2004).
A leading theory for filopodia generation proposes that these structures emerge from a rearrangement of the dendritic networks of the lamellipod (Svitkina et al., 2003). In the convergence/elongation theory, convergent filaments with barbed ends that abut the plasma and are protected against capping are zippered together by fascin as they grow to several microns in length. Consistent with this model, recent knockdown studies reveal that filopodia-rich phenotypes occur in capping-protein-depleted cells and point to an essential role for the anticapping activies of the Ena/VASP family in filopodia formation (Mejillano et al., 2004).
Nearer the cell body, filaments surviving the destructive actions of ADF/cofilins mature into a contractile network. In fast-moving cells like fish keratocytes, the surviving network remains fixed with respect to the substrate as the cell crawls past (Theriot and Mitchison, 1991). In slower-moving cells like fibroblasts, polymerization exceeds the rate of cell advancement, and much of the network flows toward the cell center (Theriot and Mitchison, 1992). Filament survival is facilitated by association with the long, side-binding protein tropomyosin, which blocks the association of ADF/cofilins (Bernstein and Bamburg, 1982; Cooper, 2002; DesMarais et al., 2002), and growth to several microns is likely facilitated by the anticapping activities of formins (Higashida et al., 2004). In fibroblasts, the polarity of filaments is graded such that all filaments at the leading edge are oriented with their barbed ends facing the periphery, but the interior bundles have a well-mixed polarity (Cramer et al., 1997). The gradation appears to facilitate both pushing at the edge of cells and myosin-based contractions by muscle-like filaments sliding within the cell interior. Just as some of the filaments of the lamellipod mature into contractile stress fibers, the focal contacts that transmit these stresses to surfaces also begin life in lamellipodia as nascent focal complexes and mature into focal contacts as they become part of the more central structures of the cell (DeMali et al., 2002).
Monomer recycling: the other ‘actin dynamics’
For steady migration a cell must have a constant supply of monomer delivered to its leading edge. This monomer certainly derives from filament disassembly at more interior regions, and so the rates of assembly and migration are tied to the rate of monomer supply. If monomer is provided by diffusion, the supply rate is equal to the product of the diffusion coefficient and the gradient of the monomer concentration. The diffusion coefficient of the fastest of two kinetically distinguishable populations of actin tracers is 6 × 10−8 cm2/s (Giuliano and Taylor, 1994; Luby-Phelps et al., 1985; McGrath et al., 1998a). Assigning this diffusion coefficient to actin monomer has caused difficulties for modelers of the lamellipod (Abraham et al., 1999; Mogilner and Edelstein-Keshet, 2002). If the assembly of monomers at the leading edge of a
192J.L. McGrath and C.F. Dewey, Jr.
crawling cell is driven by mass action, then the concentration of monomer at the leading edge must exceed 15 µM for a single filament to keep pace with the plasma
membrane in keratocytes. Fick’s law, in combination with a monomer diffusion coefficient of 6 × 10−8 cm2/s, requires a gradient of 20 µM/µm moving toward the
interior of the cell. If this gradient persists over 10 microns, as modelers have assumed, then a maximum concentration of 200 µM G-actin near the cell body is prohibitively high because cells typically carry less than 100 µM of actin total.
Assuming that the more diffusive population of actin is strictly monomer may be wrong, as short, diffusing filaments from recent severing and growth events are likely. Indeed, investigators have proposed that the diffusion of small oligomers may explain why the diffusion coefficient for the mobile actin in cytoplasm is 6 times slower than the value for monomer-sized ficoll (Luby-Phelps et al., 1987). Through comparisons with the diffusion of sugar particles of various sizes, one concludes that actin diffuses as a molecule five times bigger than its hydrodynamic radius (LubyPhelps et al., 1987), and so the discrepancy cannot be explained by the fact that G-actin is complexed with smaller molecules of thymosin or profilin. The possibility
of filament diffusion appears to justify the use of the high G-actin diffusion coefficient of 3 × 10−7 cm2/s in models (Abraham et al., 1999; Mogilner and Edelstein-Keshet,
2002). The value is that for the diffusion of monomer-sized ficoll and leads to the derivation of G-actin profiles with more gradual gradients of 4 µM/µm. However it must be noted that all proteins studied diffuse slower in cytoplasm than size-matched ‘inert’ sugars, even those that do not oligomerize (Luby-Phelps et al., 1985), and so the most reasonable explanation for the discrepancy between the diffusion of actin and sugar particles is that the sugars are not good models for protein diffusivity.
In support of the interpretation of the fast-diffusing species as monomer, roughly half of the tracer actin is in this population consistent with biochemical fractionation (McGrath et al., 2000c). Further, when cells are treated with jasplankinolide, a membrane-permeating toxin that blocks filament depolymerization (Bubb et al., 2000), all actin becomes immobile, indicating that the fast-diffusing species is assembly competent (McGrath et al., 1998a; Zicha et al., 2003). As noted, recent speckle microscopy experiments indicate that the major zone of depolymerization in lamellipodia is found 2 microns behind the leading edge (Ponti et al., 2004; Watanabe and Mitchison, 2002), and so the gradient may in fact be steep but confined to a dynamic region considerably smaller than the 10 microns assumed in the models. Over this small distance the experimentally determined values for monomer diffusion should be sufficient for the steady resupply of monomer to the leading edge.
Recent data also suggest that G-actin is returned to the leading edge of cells by active transport mechanisms. This prospect is raised by the recent results of Grahm Dunn and colleagues (Zicha et al., 2003). Using a modification of the photobleaching technique, these experimentalists marked a population of actin several microns behind the leading edge of a protruding cell and found that some of the marked actin incorporates into new protrusions at rates that defy simple diffusion. They provide some evidence that the phenomenon is halted by myosin inhibitors and does not depend on microtubules. This suggests that the privileged monomer does not ride as cargo on a motor complex, but that it happens to be near a convective channel. The difficulty with the prospect of motors moving G-actin as cargo is that the association between
