
- •Contents
- •Contributors
- •Preface
- •1 Introduction, with the biological basis for cell mechanics
- •Introduction
- •The role of cell mechanics in biological function
- •Maintenance of cell shape
- •Cell migration
- •Mechanosensing
- •Stress responses and the role of mechanical forces in disease
- •Active cell contraction
- •Structural anatomy of a cell
- •The extracellular matrix and its attachment to cells
- •Transmission of force to the cytoskeleton and the role of the lipid bilayer
- •Intracellular structures
- •Overview
- •References
- •2 Experimental measurements of intracellular mechanics
- •Introduction
- •Forces to which cells are exposed in a biological context
- •Methods to measure intracellular rheology by macrorheology, diffusion, and sedimentation
- •Whole cell aggregates
- •Sedimentation of particles
- •Diffusion
- •Mechanical indentation of the cell surface
- •Glass microneedles
- •Cell poker
- •Atomic force microscopy
- •Mechanical tension applied to the cell membrane
- •Shearing and compression between microplates
- •Optical traps
- •Magnetic methods
- •Twisting of magnetized particles on the cell surface and interior
- •Passive microrheology
- •Optically detected individual probes
- •One-particle method
- •Two-particle methods
- •Dynamic light scattering and diffusing wave spectroscopy
- •Fluorescence correlation spectroscopy
- •Optical stretcher
- •Acoustic microscopy
- •Outstanding issues and future directions
- •References
- •3 The cytoskeleton as a soft glassy material
- •Introduction
- •Magnetic Twisting Cytometry (MTC)
- •Measurements of cell mechanics
- •The structural damping equation
- •Reduction of variables
- •Universality
- •Scaling the data
- •Collapse onto master curves
- •Theory of soft glassy rheology
- •What are soft glassy materials
- •Sollich’s theory of SGMs
- •Soft glassy rheology and structural damping
- •Open questions
- •Biological insights from SGR theory
- •Malleability of airway smooth muscle
- •Conclusion
- •References
- •4 Continuum elastic or viscoelastic models for the cell
- •Introduction
- •Purpose of continuum models
- •Principles of continuum models
- •Boundary conditions
- •Mechanical and material characteristics
- •Example of studied cell types
- •Blood cells: leukocytes and erythrocytes
- •Limitations of continuum model
- •Conclusion
- •References
- •5 Multiphasic models of cell mechanics
- •Introduction
- •Biphasic poroviscoelastic models of cell mechanics
- •Analysis of cell mechanical tests
- •Micropipette aspiration
- •Cells
- •Biphasic properties of the pericellular matrix
- •Indentation studies of cell multiphasic properties
- •Analysis of cell–matrix interactions using multiphasic models
- •Summary
- •References
- •6 Models of cytoskeletal mechanics based on tensegrity
- •Introduction
- •The cellular tensegrity model
- •The cellular tensegrity model
- •Do living cells behave as predicted by the tensegrity model?
- •Circumstantial evidence
- •Prestress-induced stiffening
- •Action at a distance
- •Do microtubules carry compression?
- •Summary
- •Examples of mathematical models of the cytoskeleton based on tensegrity
- •The cortical membrane model
- •Tensed cable nets
- •Cable-and-strut model
- •Summary
- •Tensegrity and cellular dynamics
- •Conclusion
- •Acknowledgement
- •References
- •7 Cells, gels, and mechanics
- •Introduction
- •Problems with the aqueous-solution-based paradigm
- •Cells as gels
- •Cell dynamics
- •Gels and motion
- •Secretion
- •Muscle contraction
- •Conclusion
- •Acknowledgement
- •References
- •8 Polymer-based models of cytoskeletal networks
- •Introduction
- •The worm-like chain model
- •Force-extension of single chains
- •Dynamics of single chains
- •Network elasticity
- •Nonlinear response
- •Discussion
- •References
- •9 Cell dynamics and the actin cytoskeleton
- •Introduction: The role of actin in the cell
- •Interaction of the cell cytoskeleton with the outside environment
- •The role of cytoskeletal structure
- •Actin mechanics
- •Actin dynamics
- •The emergence of actin dynamics
- •The intrinsic dynamics of actin
- •Regulation of dynamics by actin-binding proteins
- •Capping protein: ‘decommissioning’ the old
- •Gelsolin: rapid remodeling in one or two steps
- •β4-thymosin: accounting (sometimes) for the other half
- •Dynamic actin in crawling cells
- •Actin in the leading edge
- •Monomer recycling: the other ‘actin dynamics’
- •The biophysics of actin-based pushing
- •Conclusion
- •Acknowledgements
- •References
- •10 Active cellular protrusion: continuum theories and models
- •Cellular protrusion: the standard cartoon
- •The RIF formalism
- •Mass conservation
- •Momentum conservation
- •Boundary conditions
- •Cytoskeletal theories of cellular protrusion
- •Network–membrane interactions
- •Network dynamics near the membrane
- •Special cases of network–membrane interaction: polymerization force, brownian and motor ratchets
- •Network–network interactions
- •Network dynamics with swelling
- •Other theories of protrusion
- •Numerical implementation of the RIF formalism
- •An example of cellular protrusion
- •Protrusion driven by membrane–cytoskeleton repulsion
- •Protrusion driven by cytoskeletal swelling
- •Discussion
- •Conclusions
- •References
- •11 Summary
- •References
- •Index
Cell dynamics and the actin cytoskeleton |
193 |
actin and motor proteins would likely be specific. The prospect of a return mechanism that does not discriminate among molecules is more attractive because the problem of recycling cytoskeletal pieces is not limited to G-actin; whatever mechanism is at work must be able to carry all the building blocks to the leading edge.
The biophysics of actin-based pushing
While compelling support for actin polymerization forces have existed for decades (Tilney et al., 1973), the mechanism by which polymerization leads to pushing remains unclear. Today there are two leading theories: (1) a series of related ‘ratchet’ models that explain pushing as a natural consequence of the polymerization of semiflexible filaments against a membrane (Dickinson and Purich, 2002; Mogilner and Oster, 1996; Mogilner and Oster, 2003; Peskin et al., 1993) and (2) a mesoscopic model that explains how pushing forces derive from the formation of actin networks on curved surfaces.
Understanding of the biophysics of actin-based pushing in the Listeria system has progressed through a steadily tightening cycle of theory and experiment that continues to this day. In large part, these efforts have centered around the actin-based motility of the bacterium Listeria monocytogenes. This intracellular pathogen invades host cytoplasm and hi-jacks the same force-producing mechanisms that drive leadingedge motility. Riding a wave of actin polymerization, the bacterium becomes motile so that it can eventually exit the dying infected cell for an uninfected neighbor.
The first ratchet model proposed that Listeria was a 1-D Brownian particle blocked from rearward diffusion by the presence of the growing actin tail (Mogilner and Oster, 1996). Observations that Shigella move at speeds similar to Listeria despite being more than twice as large violated a prediction of the Brownian Ratchet and motivated a new theory. The “Elastic Ratchet” proposed that filaments, rather than Listeria, fluctuate due to thermal excitation (Mogilner and Oster, 1996). Filaments fluctuate away from the Listeria surface to allow space for polymerization. Lengthened filaments apply propulsive pressure as they relax to unstrained configurations.
More recent biophysical measurements established that Listeria are tightly bound to their tails (Gerbal et al., 2000a; Kuo and McGrath, 2000), rigorously eliminating the Brownian Ratchet theory and demanding a revision of the Elastic Ratchet theory. One study also established that Listeria motion is discontinuous, with frequent pauses and nanometer-sized steps (Kuo and McGrath, 2000). The developments led to the first proposal for how elastic filaments could push Listeria while attached. In the ‘Actoclampin’ model (Dickinson and Purich, 2002), filaments diffuse axially due to bending fluctuations within a surface-bound complex. The complex binds ATP-bound subunits and releases the filament upon hydrolysis. In this scheme, flexed filaments push the Listeria and lagging filaments act as tethers (Fig. 9-10A).
More than molecular stepping, the force-velocity curve of the polymerization engine constrains theoretical models, but current data appear to be in disagreement. Recently, we published a curve for Listeria monocytogenes (McGrath et al., 2003), using methylcellulose to manipulate the viscoelasticity of extracts, particle tracking to determine viscoelastic parameters near motile Listeria, and a modified Stokes equation to infer forces. In 2003, Mogilner and Oster published an evolution of the
194 J.L. McGrath and C.F. Dewey, Jr.
FL |
fw |
fa |
|
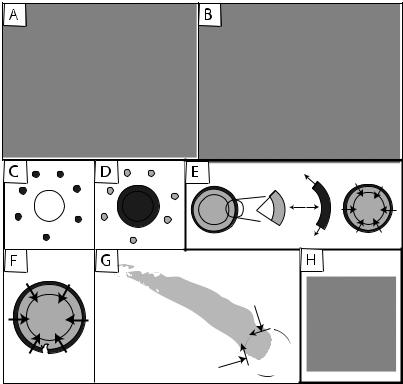
Fig. 9-10. Theories of force production in actin-based motility. (A–B) In Molecular Ratchet models, filament fluctuations and growth near the surface combine to create motility. (A) In the Actoclampin model (from Dickinson and Purich, 2002) all filaments are similarly attached to the motile surface, but some are compressed and others are stretched taut. (B) The Tethered Ratchet (from Mogilner and Oster, 2003), considers distinct attached and pushing (working) filaments. (C–G) In the Elastic Propulsion theory, elastic stresses lead to symmetry breaking and motion. (C) The first layer of actin polymerized at the surface is pushed outward (D) by the next layer, creating hoop stresses in the gel and normal pressure on the sphere (E). Fluctuations in stress levels and strengths cause a local fracturing event (F) and unraveling of the gel that leads to a motile state (G) in which stresses build and relax periodically as the particle moves forward. (H) Large particles create tails with periodic actin density (‘hopping’), suggesting stress building and relaxation. Bar is 10 microns. From Bernheim-Groswasser, 2003.
Elastic Ratchet model (Mogilner and Oster, 2003) that quantitatively predicts the force-velocity curve of McGrath et al. (2003). The “Tethered Ratchet” features “working” filaments that push as Elastic Ratchets and “tethered” filaments anchored at complexes that also nucleate dendritic branches (Fig. 9-10B).
While Molecular Ratchets appear to account for the force-velocity data in McGrath et al. (2003), shallower force-velocity curves obtained by Wiesner et al. (2003) are interpreted in terms of a very different theory termed Elastic Propulsion (Gerbal et al., 2000b) (Fig. 9-10C–H). The theory describes stress build-up in continuum, elastic actin networks that grow on curved surfaces. During nucleation, older layers are displaced radially by newer polymerization at the nucleating surface. Because the displaced layers must stretch, they generate ‘hoop’ or ‘squeezing’ stresses around the
