
Micro-Nano Technology for Genomics and Proteomics BioMEMs - Ozkan
.pdf
142 |
MICHAEL J. HELLER, CENGIZ S. OZKAN AND MIHRIMAH OZKAN |
FIGURE 6.5. A more advanced version of Nanogen’s microelectronic DNA chip device with 100 test-sites or microlocations. The test sites are approximately 80 microns in diameter, with underlying platinum microelectrodes. The outside ring of twenty micro-electrodes can be used as counter electrodes for the test-sites.
been commercialized, has an inner set of 80-micron diameter test sites with underlying platinum microelectrodes, and an outside set of auxiliary microelectrodes (Figure 6.5). The outer group of electrodes can be negatively biased, which allows DNA in the bulk sample solution to be concentrated at the specific internal test-sites (positively biased). Each microelectrode has an individual wire interconnect through which current and voltage are applied and regulated. The 100 test-site DNA chip is about 7 millimeters square in size, with an active test-site array area of about 2 millimeters. The chips are fabricated from silicon wafers, with insulating layers of silicon dioxide, platinum microelectrodes and gold connecting wires. Silicon dioxide/silicon nitride is used to cover and insulate the conducting wires, but not the surface of the platinum microelectrodes. The whole surface of the array is covered with several microns of hydrogel (agarose or polyacrylamide) which forms a permeation layer. The permeation layer is impregnated with a coupling agent (streptavidin) which allows attachment of biotinylated DNA probes or other entities [12–14, 17]. The ability to use silicon and microlithrography for fabrication of the DNA chips allows a wide variety of devices to be designed and tested (Figure 6.4). The higher density arrays (400 to 10,000) represent more sophisticated devices that have on-chip CMOS control elements for regulating the current and voltages to the microelectrode at each test site [26]. These control elements are located in the underlying silicon structure and are not exposed to the aqueous

USE OF ELECTRIC FIELD ARRAY DEVICES FOR ASSISTED ASSEMBLY |
143 |
FIGURE 6.6. Shows the NanoChipTM Molecular Biology Workstation for bioresearch and DNA diagnostic applications. This system provides a chip loader component, fluorescent detection/reader component, computer control interface and data display component. The probe loader component is used for automated DNA probe addressing or spotting, and provides the end-user with “Make Your Own Chip” capabilities. Insert shows the NanoChipTM cartridge device. A 100 test site microelectronic array incorporated into a cartridge package that provides the electronic, optical, and fluidic interfacing.
samples that are applied to the chip surface when carrying out the DNA hybridization reactions.
The 100 test-site microelectronic array device has been incorporated into a cartridge package (NanoChipTM cartridge) which provides for the electronic, optical, and fluidic interfacing. The NanoChipTM cartridge assembly is shown as the insert in Figure 6.6. The chip itself is mounted (flip chip bonded) onto a ceramic plate and pinned out for electrical connections. The chip/ceramic plate component is mounted into a plastic cartridge that provides several fluidic input and output ports for addition and removal of DNA samples and reagents. The area over the active test-site portion of the array is an enclosed sample chamber covered with a quartz glass window. This window allows for fluorescent detection to be carried out on the hybridization reactions that occur at the test sites on the array surface. A complete instrument system (NanoChipTM Molecular Biology Workstation) provides a chip loader component, a fluorescent detection/reader component, a computer control interface and a data display screen component (Figure 6.6). The probe loading component allows DNA probes or target sequences (DNA, RNA, PCR amplicons) to be selectively addressed to the array test-sites, and provides a “make your own chip” capability. The automated probe loader system allows four 100 test-site NanoChipTM cartridges to be loaded

144 |
MICHAEL J. HELLER, CENGIZ S. OZKAN AND MIHRIMAH OZKAN |
FIGURE 6.7. Demonstration of checker-boarding of fluorescent DNA molecules on a microelectronic array. Checker-boarding is achieved by DC biasing alternate test sites on the array positive and negative, and then switching the DC electric field bias every 6 seconds.
with DNA probes or samples from a ninety-six or three hundred eighty four well microtiter plates. Probes or target sequences are usually biotinylated, which allows them to become bound to streptavidin within the permeation layer of the specified test-site. In the electronic addressing procedure the probe loader component deliverers the desired biotinylated probe to specified test sites which are biased positive. The electric (electrophoretic) field causes the negatively charged DNA molecules to concentrate onto the positively activated test-sites, with subsequent binding via the biotin/streptavidin reaction. The ability of the electric field to concentrate DNA molecules on to specific test sites on the array is shown in Figure 6.7. The figure shows the controlled parallel movement (checker-boarding) of negatively charged fluorescent DNA molecules between alternating positively and negatively biased microlocations. The sample solution contains about 50 nM concentration of a 20-mer oligonucleotide sequence which is labeled in the 5’-terminal position with Bodipy Texas Red fluorophore (Ex 590 nm, Em 620 nm), the microelectrodes are biased positive and negative in a checkerboard fashion, and the field is reversed every six seconds. At about 3 volts DC the fluorescent DNA molecules ( 7 nm in length) are transported back and forth a distance of 200 microns during the six second switching time. In actuality, when DNA hybridization reactions are carried out on these active microelectronic arrays, controlled DC electric fields are first used to transport and address (spot) specific DNA molecules to the selected test-sites/microlocations on the array. Such oligonucleotide “capture” probes or target DNA sequences are usually functionalized with biotin molecules, and become strongly bound to streptavidin molecules which are cross-linked to the hydrogel layer covering the underlying microelectrode. In the next steps, electric fields are used to control and direct the hybridization of the other DNA molecules to the DNA sequences attached to the selected test sites. Electronic addressing also allows DNA probes to be spotted onto the array in a highly reproducible manner. Microelectronic arrays can be formatted in a variety of ways that include reverse dot blot format (capture/ identity sequences bound to test-sites), sandwich format (capture sequences bound to test-sites) and dot blot format (target sequences bound to test-sites). DNA hybridization assays involve the use of fluorescent reporter probes or target DNA sequences. The reporter groups are usually organic fluorophores that have been either attached to oligonucleotide probes or to the target/sample DNA/RNA sequences. After electronic addressing and hybridization are carried out, the chip is analyzed using the fluorescent detection system. The fluorescent detector has two different laser excitation sources (excitation 532 nm and excitation 635 nm). The laser beams are quickly scanned across the array using a confocal type optical system and the emissions from the fluorescent labeled probes or targets (550–600 nm “green” and 660–720 nm
USE OF ELECTRIC FIELD ARRAY DEVICES FOR ASSISTED ASSEMBLY |
145 |
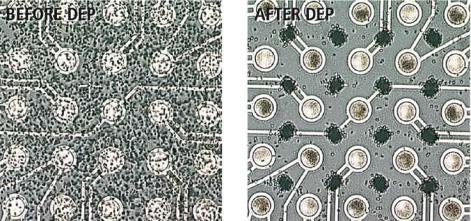
FIGURE 6.8. Shows the separation of Listeria from whole blood by DEP on a microelectronic array (Before and After DEP). Listeria cells become positioned on the high AC field (the microlocations), and the blood cells become aggregated in the low AC field regions between the microlocations.
“red”) are detected. References [27–36] provide some examples of single nucleotide polymorphism (SNP) genotyping analysis that is carried out on these microelectronic arrays. For genotyping analysis, the selection rules are simple and no complex image processing is required for data analysis, match/mismatch calls are easy to make and results are highly reliable.
Microelectronic arrays can also be used for a variety of cell separation applications. Disease diagnostics frequently involve identifying a small number of specific bacteria or viruses in a blood sample (infectious disease), fetal cells in maternal blood (genetic diseases) or tumor cells in a background of normal cells (early cancer detection). While a number of techniques are available, electric field based methods have been developed and which can have considerable advantages. One basic electronic method for cell separation is called dielectrophoresis (DEP). This process involves the application of an asymmetric alternating current (AC) electric field to the cell population. Active microelectronic arrays have been used to achieve the separation of bacteria from whole blood [37], and for the separation of cervical carcinoma cells from blood [38] and for gene expression analysis [39]. Figure 6.8 shows a microelectronic array device utilizing high frequency AC fields to carry out the DEP separation of Listeria bacterial cells ( 1 µm) from whole blood cells ( 10 µm) in a highly parallel manner. At an AC frequency of about 10 kHz the Listeria bacterial cells are positioned on the microlocations (high field region), and the blood cells are positioned in the low field regions between the microelectrodes. The relative positioning of the cells between the high and low field regions is based on dielectric differences between the cell types. In Figure 6.8, the left panel shows microelectronic array with the mixed blood/bacteria sample, and the right panel shows the cell separation pattern which occurs after a high frequency AC electric field is applied. While maintaining the AC field, the microarray can be washed with a buffer solution that removes the blood cells (low field regions) from the more firmly bound bacteria (high field regions) near the microelectrodes. The bacteria can then be released and collected or electronically lysed to release the genomic DNA or RNA
146 |
MICHAEL J. HELLER, CENGIZ S. OZKAN AND MIHRIMAH OZKAN |
for further manipulation and analysis [37]. DEP represents a particularly useful process that allows difficult cell separation applications to be carried out rapidly and with high selectivity. The DEP process can also be extremely useful for nanofabrication purposes.
6.3. ELECTRIC FIELD ASSISTED NANOFABRICATION PROCESS
Enabling Nanotechnology—Generally, molecular or nanoelectronic devices and systems are envisioned as the more revolutionary application of this new technology. Presently, many examples of individual molecular components with appropriate basic properties exist and include entities such as carbon nanotubes, nanowires and various organic molecules with electronic switching capabilities. However, the larger issue with enabling molecular electronics is more likely to be the development of a viable technology which will allow billions of molecular or nanoelectronic components to be assembled and interconnected into useful logic/memory devices and systems. In addition to the electronic applications, nanodevices and nanosystems with higher order photonic, mechanical, mechanistic, sensory, chemical, catalytic, and therapeutic properties are also envisioned [1–4]. Again, key problems with enabling such devices and systems will most likely occur at the stage of organizing components for higher level functioning, rather than the availability of the molecular components. By way of example, Figure 6.1 shows a comparison between a biological photonic energy conversion system, “a plant chloroplast”, and a man-made charged coupled device (CCD). The chloroplast is made by a self-assembly “bottom-up process, while the CCD device is made by microlithography, a top-down process. In the chloroplast, hundreds of relatively simple chromophore molecules are arranged into solid state antenna structures which collect photonic energy and transfer it through the structure with high quantum efficiency. The plant chloroplast (a few microns in size) represents a highly integrated light-capturing device composed of numerous self-organized molecular and nanoscale structures. In the man-made CCD device the feature size stops at about the micron-scale level. In addition to the chloroplast being composed of nanoscale structures, the energy transfer mechanism used within the structure is also very unique. The transfer of photonic energy through the antenna structures is carried out by what is called the Forster resonant energy transfer mechanism [40]. In some sense, this process might be thought of as the equivalent of molecular fiber optics, but it is quite different than any man-made process or device for photonic energy collection and transfer. Such a mechanism could potentially be very useful for communication between molecular and nanoscale components and devices. To date, it has been difficult to design a synthetic model of this type of solid state photonic transfer structure with any of the properties or efficiencies seen in the biological systems. Thus, a key problem with mimicking this type of bionanotechnology is the lack of a suitable “bottom-up” fabrication process to carry out the precision integration of diverse molecular and nanoscale components into viable higher order structures.
Electric Field Array Approach to Nanofabrication—As was stated earlier, microelectronic array devices have been developed for applications in genomic research and DNA diagnostics. These active microelectronic devices have the ability to create a variety of re-configurable electric field geometries on the surface of the array device. This capability allows almost any type of charged molecule or structure (DNA, RNA, protein, enzyme, nanostructure, cell, or micron-scale device) to be electrophoretically transported to or from
USE OF ELECTRIC FIELD ARRAY DEVICES FOR ASSISTED ASSEMBLY |
147 |
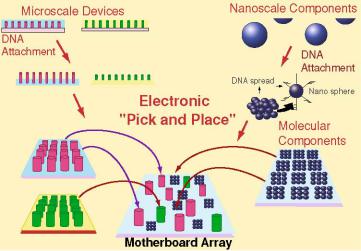
FIGURE 6.9. An electric field based “pick and place” process which uses a microelectronic “Mother Board” array to carry out the heterogeneous integration of molecular, nanoscale and microscale components into higher order devices and structures.
any of the microscopic sites (microlocations). When specific DNA hybridization reactions are carried out on this type of active microarray, the device is actually using electric fields to direct the self-assembly of DNA molecules at the specified test-site or microlocation on the chip surface. Thus, in principle these active devices serve as a “mother-board or host board” for the assisted assembly of DNA molecules into more complex structures. Since the DNA molecule with its intrinsic programmable self-assembly properties can be derivatized with electronic or photonic groups or attached to larger nanostructures (quantum dots, metallic nanoparticles, nanotubes), or microstructures, we have a unique process for nanofabrication. In essence, these active microelectronic mother-board arrays allow one to carry out a highly parallel electric field “pick & place” process for the heterogeneous integration of molecular, nanoscale and micron-scale components into complex three dimensional structures within the defined perimeters of larger silicon or semiconductor structures (Figure 6.9). The electric field assisted self-assembly technology is based on three key physical principles:
(1) the use of functionalized DNA structures as “Lego” blocks for nanofabrication; (2) the use of DNA as a selective “glue” that provides intrinsic self-assembly properties to molecular and nanoscale electronic components and structures (carbon nanotubes, organic molecular electronic switches, etc.); and (3) the use of an active microelectronic array for electric field assisted self-assembly of any modified electronic/photonic components and structures into integrated structures [23–25]. Microelectronic arrays which are produced by the “topdown” process of photolithography, thus have an intrinsic capability to direct the assembly of molecular structures via a “bottom-up” process.
Microelectronic arrays have been used to direct the binding of derivatized nanospheres and microspheres on to selected locations on the microarray surface. In this case, fluorescent and non-fluorescent polystyrene nanospheres and microspheres derivatized with specific DNA oligonucleotides are transported and bound to selected test sites or microlocations

148 |
MICHAEL J. HELLER, CENGIZ S. OZKAN AND MIHRIMAH OZKAN |
|
A |
B |
C |
|
||
D |
E |
|
|
|
1 m |
100 nm |
10 m 2 m |
|
5 m |
FIGURE 6.10. Demonstrates the selective addressing of 10, 5, 2 and 1 micron microspheres and 100 nanometer nanospheres (red arrows) using a 25 test site microelectronic array device. A. shows array before spheres are added. B, C, show stages in addition of the various spheres; D shows all spheres addressed, and E show all sphere spheres in dark field (nanospheres now visible).
derivatized with the specific complementary oligonucleotide sequences. Thus, microelectronic array devices are not just limited to selective transport and binding of small molecules such as fluorescent DNA, but also for selective transport and addressing of larger nanoparticles and microspheres, and even objects as large as 20 micron light emitting diode structures [21, 22, 24]. By way of example, Figure 6.10 shows a sequence of frames demonstrating the addressing of derivatized negatively charged polystyrene microspheres and nanospheres to selectively activated microlocations on a 25 test site microelectronic array. The addressing sequence is first, the one micron microspheres, second the two micron microspheres, third the 100 nanometer nanospheres, fourth the five micron microspheres and finally the ten micron microspheres. The addressing time for each group of spheres is about 10 to 15 seconds at about 3 volts DC. The final figure in the sequence shows a dark field contrast micrograph which allows the 100 nm nanospheres packed on the 80 µm diameter microlocation to be better visualized. Thus, microelectronic array devices can serve as “motherboards or hostboards” for the directed assembly of more complex nanostructures and microscale structures on their surface. A further advantage of the active microarray devices is that they can be operated in both DC and AC modes. In the DC electric field mode the electrophoretic transport of charged molecules and nanostructures is carried out between the positive and negative biased microlocations. The rate of transport is related to the strength of the electric field and the charge/mass ratio of the molecule or structure. When the microelectronic arrays are used to generate high frequency AC fields they can carry out the process of dielectrophoresis (DEP). With high AC fields, nanostructures, cells and other micron scale structures can be oriented and selectively positioned based on their
USE OF ELECTRIC FIELD ARRAY DEVICES FOR ASSISTED ASSEMBLY |
149 |
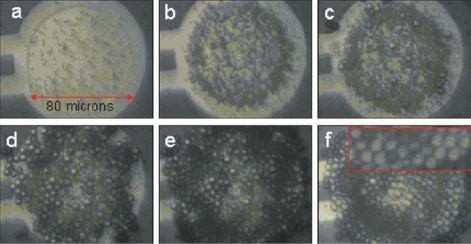
FIGURE 6.11. Shows the electric field addressing of derivatized negatively charged one micron size polystyrene microspheres to a selectively activated microlocation on a 25 test site microelectronic array, and then the addressing of the larger five micron microspheres over the one micron microspheres. Red box in (f) shows a magnified view of the packed five micron microsphere layer.
intrinsic dielectric properties. Figure 6.11 shows a sequence of frames demonstrating the addressing of derivatized negatively charged one micron size polystyrene microspheres to a selectively activated microlocation on a 25-site microelectronic array. After the microlocation is layered with the one micron polystyrene microspheres, a second addressing sequence is carried out in which larger five micron microspheres are now layered over the one micron sized polystyrene microspheres. Thus, it is possible to use electric field addressing to layer particles and other materials, allowing fabrication in the third dimension. In addition to DNA assisted assembly, other chemical and biological entities can be used advantageously for device forming, pick and place, as well as for micro and nanoactuation.
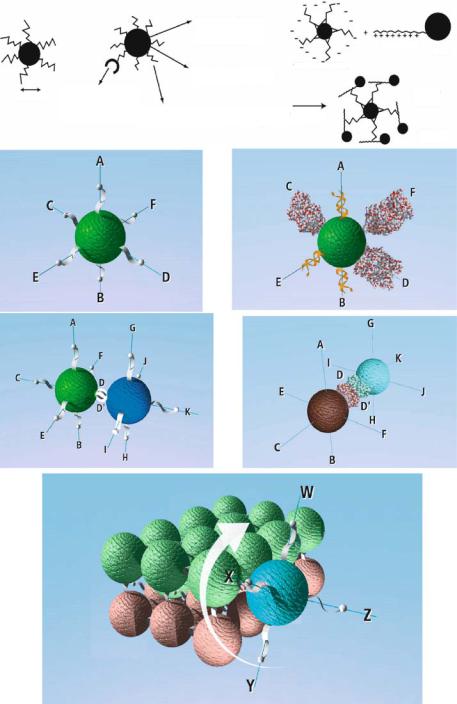
Present nanofabrication methods do not allow most nanostructures to be modified in a controlled or precise manner. For example, Figure 6.12 (upper section) shows a present scheme for modifying a nanoparticle with DNA or protein ligands using classical chemical functionalization procedures. While such a process allows a higher-order structure to be formed, there is little or no control as to how the DNA or protein ligands are arranged around the core nanostructure. Furthermore, it is not a viable process for arranging “different” DNA, protein or binding entities around the core nanostructure in a defined fashion. Microelectronic array devices may offer the opportunity to development processes that will allow core nanostructures to be selectively modified as show in Figures 6.12a & b. In this case, the nanostructures are derivatized at specific positions around the core with specific DNA sequences (oligonucleotides) proteins, or other ligands. Such modifications allow a given nanostructure to be joined with another nanostructure in a more precise manner, i.e., leaving other selected positions on the nanostructure available for binding different nanostructures (Figure 6.12c & d). The ability to produce precision nanocomponents with selective ligands as shown above is will be necessary in order to carry out further organization into more complex “heterogeneous” three dimensional structures. The need for precision placement
|
Functional Groups: |
1-30 nm |
"Chemical Hooks or other |
|
chemically, electrically, |
|
optically active groups" |
Core : "Active Material" 1-30 nm Size Spheres, Rods, Disks, etc...
Shells: "Proctive or
Complementary Layers"
Surface Groups: "Passivating, Proctective, and
Chemically Active Layer"
"+" charged zipper
Protein
QD decorated with "-" charges
Quantum Dot Decorated with Proteins
Electrostatic Self-
Assembly
Mattoussi et al, (IACS, 2000)
FIGURE 6.12. Upper figure section shows a classical scheme for the non-precision functionalization of nanospheres. Sections a, b, c & d shows a precision nanosphere functionalization scheme; with c&d showing the selective binding of two precision nanospheres. Final figure shows the type of heterogeneous 3D higher order structure that can only be obtained using precision nanostructures.
USE OF ELECTRIC FIELD ARRAY DEVICES FOR ASSISTED ASSEMBLY |
151 |
of ligands on nanostructures becomes even more important when one observes that even a small amount of rotation around the axis of one ligand may disrupt the further formation of a higher order 3D structure (Figure 6.12d).
The proposed microelectronic array techniques have the capability to carry out the precision functionalization of nanostructures by processes which involve transporting and orienting the nanostructures onto surfaces containing the selected ligand molecules which are then reacted only with a selected portion of the nanostructure. By repeating the process and reorienting the nanostructures it will be possible to functionalize the core structure selectively with most biological and/or chemical groups. Such devices and processes allow one to design and create functionalized nanostructures with binding groups arranged in tetrahedral, hexagonal or other coordinate positions around the core nanostructure.
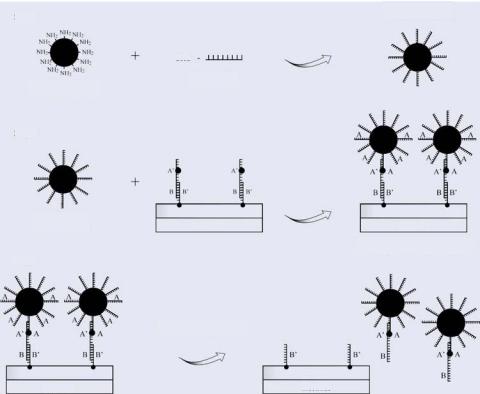
Microelectronic array devices may have the potential to carry out an electric field orientation synthesis process for producing precision nanostructures or microstructures (e.g., quantum dots, nanospheres, nanoparticles, nanocrystals) with multiple DNA identities [23– 24]. These multiple identities can be in the form of specific DNA sequences which are located at different coordinates on the particle surface. These coordinates can be tetrahedral or hexagonal in their nature. Figure 6.13 shows the initial steps for producing a precision
Step 1 |
Oligo (A) bound |
to entire surface area |
OHC
Oligo (A) with Nanosphere with aldehyde
primary amine groups
Step 2
Nanosphere as |
Electrode |
Electrode |
prepared in step 1 |
Nanospheres electronically addressed and oligos (A/A') UV crosslinked
Step 3
|
Nanosphere-oligo conjugate |
|
in de-hybridized from array |
Electrode |
Electrode |
FIGURE 6.13. Shows the first part of a scheme for the electric field based precision functionalization of nanospheres with DNA oligonucleotide sequences [24].
