
- •Preface
- •Contents
- •1 Extracellular and Intracellular Signaling – a New Approach to Diseases and Treatments
- •1.1 Introduction
- •1.1.1 Linear Model of Drug Receptor Interactions
- •1.1.2 Matrix Model of Drug Receptor Interactions
- •1.2 Experimental Approaches to Disease Treatment
- •1.3 Adipokines and Disease Causation
- •1.4 Questions in Disease Treatment
- •1.5 Toxic Lifestyles and Disease Treatment
- •References
- •2.1 Introduction
- •2.2 Heterogeneity of Adipose Tissue Composition in Relation to Adipokine and Cytokine Secretion
- •2.3 Feedback between FA and the Adipocyte
- •2.6 Metabolic Programming of Autocrine Signaling in Adipose Tissue
- •2.8 Cell Heterogeneity in the Pancreatic Islet
- •2.16 Concluding Remarks
- •Acknowledgements
- •References
- •3 One Receptor for Multiple Pathways: Focus on Leptin Signaling
- •3.1 Leptin
- •3.2 Leptin Receptors
- •3.3 Leptin Receptor Signaling
- •3.3.4 AMPK
- •3.3.5 SOCS3
- •3.4 Leptin Receptor Interactions
- •3.4.1 Apolipoprotein D
- •3.4.2 Sorting Nexin Molecules
- •3.4.3 Diacylglycerol Kinase Zeta
- •3.4.4 Apolipoprotein J
- •References
- •4.1 Introduction
- •4.2 Leptin: A Brief Introduction
- •4.3 Expression of Leptin Receptors in Cardiovascular Tissues
- •4.6 Post Receptor Leptin Signaling
- •4.6.2 Mitogen Activated Protein Kinase Stimulation
- •4.7 Adiponectin
- •4.7.1 Adiponectin and Cardiovascular Disease
- •4.7.2 Adiponectin and Experimental Cardiac Hypertrophy
- •4.8 Resistin
- •4.8.1 Cardiac Actions of Resistin
- •4.8.1.1 Experimental Studies on the Cardiac Actions of Resistin
- •4.9 Apelin
- •4.9.1 Apelin and Heart Disease
- •4.10 Visfatin
- •4.11 Other Novel Adipokines
- •4.12 Summary, Conclusions and Future Directions
- •Acknowledgements
- •References
- •5 Regulation of Muscle Proteostasis via Extramuscular Signals
- •5.1 Basic Protein Synthesis
- •5.2.1 Hormones
- •5.2.1.1 Mechanisms of Action: Glucocorticoids
- •5.2.1.2 Mechanisms of Action: TH (T3)
- •5.2.1.3 Mechanisms of Action: Testosterone
- •5.2.1.4 Mechanisms of Action: Epinephrine
- •5.2.2 Local Factors (Autocrine/Paracrine)
- •5.2.2.1 Mechanisms of Action: Insulin/IGF Spliceoforms
- •5.2.2.2 Mechanisms of Action: Fibroblast Growth Factor (FGF)
- •5.2.2.3 Mechanisms of Action: Myostatin
- •5.2.2.4 Mechanisms of Action: Cytokines
- •5.2.2.5 Mechanisms of Action: Neurotrophins
- •5.2.2.7 Mechanisms of Action: Extracellular Matrix
- •5.2.2.8 Mechanisms of Action: Amino Acids (AA)
- •5.3 Regulation of Muscle Proteostasis in Humans
- •5.3.1 Nutrients as Regulators of Muscle Proteostasis in Man
- •5.3.2 Muscular Activity (i.e. Exercise) as a Regulator of Muscle Proteostasis
- •5.4 Conditions Associated with Alterations in Muscle Proteostasis in Humans
- •5.4.2 Disuse Atrophy
- •5.4.3 Sepsis
- •5.4.4 Burns
- •5.4.5 Cancer Cachexia
- •References
- •6 Contact Normalization: Mechanisms and Pathways to Biomarkers and Chemotherapeutic Targets
- •6.1 Introduction
- •6.2 Contact Normalization
- •6.3 Cadherins
- •6.4 Gap Junctions
- •6.5 Contact Normalization and Tumor Suppressors
- •6.6 Contact Normalization and Tumor Promoters
- •6.7 Conclusions
- •References
- •7.1 Introduction
- •7.2 Background on Migraine Headache
- •7.3 Migraine and Neuropathic Pain
- •7.4 Role of Astrocytes in Pain
- •7.5 Adipokines and Related Extracellular Signalling
- •7.6 The Future of Signaling Research to Migraine
- •Acknowledgements
- •References
- •8.1 Alzheimer’s Disease
- •8.1.2 Target for AD Therapy
- •8.2 AD and Metabolic Dysfunction
- •8.2.1 Impaired Glucose Metabolism
- •8.2.2 Lipid Disorders
- •8.2.3 Obesity
- •8.3 Adipokines
- •8.3.1 Leptin
- •8.3.2 Adiponectin
- •8.3.3 Resistin
- •8.3.4 Visfatin
- •8.3.5 Plasminogen Activator Inhibitor
- •8.3.6 Interleukin-6
- •8.4 Conclusions
- •References
- •9.1 Introduction
- •9.1.1 Structure and Function of Astrocytes
- •9.1.1.1 Morphology
- •9.1.1.2 Astrocyte Functions
- •9.1.2 Responses of Astrocytes to Injury
- •9.1.2.1 Reactive Astrocytosis
- •9.1.2.2 Cell Swelling
- •9.1.2.3 Alzheimer Type II Astrocytosis
- •9.2 Intracellular Signaling System in Reactive Astrocytes
- •9.2.1 Oxidative/Nitrosative Stress (ONS)
- •9.2.2 Protein Kinase C (PKC)
- •9.2.5 Signal Transducer and Activator of Transcription 3 (STAT3)
- •9.3 Signaling Systems in Astrocyte Swelling
- •9.3.1 Oxidative/Nitrosative Stress (ONS)
- •9.3.2 Cytokines
- •9.3.3 Protein Kinase C (PKC)
- •9.3.5 Protein Kinase G (PKG)
- •9.3.7 Signal Transducer and Activator of Transcription 3 (STAT3)
- •9.3.10 Ion Channels/Transporters/Exchangers
- •9.4 Conclusions and Perspectives
- •Acknowledgements
- •References
- •10.1 Adipokines, Toxic Lipids and the Aging Brain
- •10.1.1 Toxic Lifestyles, Adipokines and Toxic Lipids
- •10.1.2 Ceramide Toxicity in the Brain
- •10.3 Oxygen Radicals, Hydrogen Peroxide and Cell Death
- •10.4 Gene Transcription and DNA Damage
- •10.5 Conclusions
- •References
- •11.1 Introduction
- •11.2 Cellular Signaling
- •11.2.1 Types of Signaling
- •11.2.2 Membrane Proteins in Signaling
- •11.3 G Protein-Coupled Receptors
- •11.3.1 Structure of GPCRs
- •11.3.1.1 Structure Determination
- •11.3.1.2 Structural Diversity of Current GPCR Structures
- •11.3.1.3 Prediction of GPCR Structure and Ligand Binding
- •11.3.2 GPCR Activation: Conformation Driven Functional Selectivity
- •11.3.2.2 Ligand or Mutation Stabilized Ensemble of GPCR Conformations
- •11.3.2.4 GPCR Dimers and Interaction with Other Proteins
- •11.3.3 Functional Control of GPCRs by Ligands
- •11.3.3.1 Biased Agonism
- •11.3.3.2 Allosteric Ligands and Signal Modulation
- •11.3.4 Challenges in GPCR Targeted Drug Design
- •11.4 Summary and Looking Ahead
- •Acknowledgements
- •References
- •12.1 Introduction
- •12.5.1 Anthocyanins
- •12.5.2 Gallates
- •12.5.3 Quercetin
- •12.5.5 Piperine
- •12.5.6 Gingerol
- •12.5.7 Curcumin
- •12.5.8 Guggulsterone
- •12.6.1 Phytanic Acid
- •12.6.2 Dehydroabietic Acid
- •12.6.3 Geraniol
- •12.7 Agonists of LXR that Reciprocally Inhibit NF-jB
- •12.7.1 Stigmasterol
- •12.7.3 Ergosterol
- •12.8 Conclusion
- •References
- •13.1 Introduction
- •13.2 Selective Dopaminergic Neuronal Death
- •13.3 Signaling Pathways Involved in Selective Dopaminergic Neuronal Death
- •13.3.1 Initiators and Signaling Molecules
- •13.3.1.1 Response to Oxidative and Nitrosative Stress
- •13.3.1.2 Response to Altered Proteostasis
- •13.3.1.3 Response to Glutamate
- •13.3.1.4 Other Initiators
- •13.3.2 Signal Transducers, Intracellular Messengers and Upstream Elements
- •13.3.2.2 Small GTPases
- •13.3.3 Intracellular Signaling Cascades
- •13.3.3.1 Mitogen Activated Protein Kinases (MAPK) Pathway
- •13.3.3.2 PI3K/Akt Pathway
- •13.3.3.4 Unfolded Protein Response (UPR)
- •13.3.4 Potentially Involved Intracellular Signaling Components
- •13.3.4.3 PINK1
- •13.3.5.2 Dopamine Metabolism
- •13.3.5.3 Cell Cycle
- •13.3.5.4 Autophagy
- •13.3.5.5 Apoptosis
- •13.4 Conclusions
- •References
- •Subject Index

DNA, Nuclear Cell Signaling and Neurodegeneration |
179 |
membrane-bound enzyme that transfers electrons from extracellular NADH to extracellular oxygen, making extracellular superoxide.50 NADH oxidase is the major source of superoxide in endothelial cell preparations.50 Amyloidb has NADH oxidase activity and forms extracellular oxygen radicals from extracellular NADH.58 Therefore, NADH generated in the blood of aging patients may enter into a vicious cycle with the generation of oxygen radicals that may damage the vasculature and neurons.
|
|
|
|
Xanthine |
|
|
Visfatin |
CD38 |
|
dehydrogenase |
|
Nam + ATP |
NMN |
NAD |
NADH (10.1) |
||
|
|||||
|
|
ATP |
|
NADH |
|
|
|
|
|
oxidase |
|
|
|
|
Superoxide |
||
Nicotinamide (Nam) has significant pharmacological activity that is distinct from niacin.59 Nam is a powerful neuroprotective agent that has been in use in clinics to save the lives of thousands of patients with pellagra-induced dementia and brain damage since the 1930s.59 Nam is a precursor for NAD and an inhibitor of poly(ADP-ribose) polymerase (PARP). NAD is primarily involved in the repair of DNA damage in the nucleus since it is a substrate for PARP.59 Of course, NAD and NADH are involved in cellular energy metabolism, especially in mitochondria. The pharmacology of NMN is poorly described. Several compounds similar to NMN, such as cyclic ADP-ribose, ADP-ribose and NaADP (nicotinic acid ADP), are calcium-mobilizing agents.60 NMN can inhibit the enzymatic degradation of NAD by glycohydrolase.61
Visfatin is secreted by visceral adipocytes and macrophages.62 Therefore, damage to the blood-brain barrier by ceramide may attract monocytes/ macrophages that locally produce visfatin. Visfatin induces the formation of TNFa, IL-6 and other cytokines in monocytes and other cells.63 TNFa
stimulates the secretion of adhesion molecules that cause the chemotaxis of white blood cells, such as more monocytes.64–66 Il-6 stimulates the activation of monocytes.64–66 Visfatin then establishes a redox cycle that depends on
extracellular NADH and results in much more damage to the blood-brain barrier. It may be that ceramide and visfatin work together to damage the blood-brain barrier and attract macrophages.67
10.3Oxygen Radicals, Hydrogen Peroxide and Cell Death
Ceramide and visfatin appear to induce the formation of oxygen radicals including peroxynitrite and hydrogen peroxide. Peroxynitrite is charged and does not normally cross cell membranes. It is a powerful protein-nitrating agent that nitrates protein phosphatase type 2A.68 This nitration inhibits the normal
180 |
Chapter 10 |
phosphorylation that controls the enzyme. The nitrated enzyme, in endothelial cells, is dysfunctional and causes the blood-brain barrier to leak.
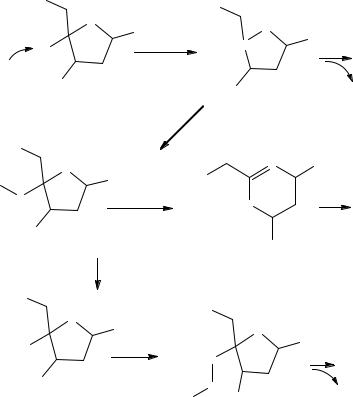
Hydrogen peroxide quickly passes through membranes and within minutes causes DNA to fragment (Figure 10.1). Hydrogen peroxide may form a hydroxyl radical that rapidly breaks DNA. DNA fragmentation occurs through three mechanisms: cleavage of the deoxyribose radical, Criegee rearrangement and peroxide migration. Radical cleavage involves cleavage of a negatively charged phosphate moiety leaving a positively charged deoxyribose radical. Criegee rearrangement involves hydroxide attack of the deoxyribose with elimination of a negatively charged phosphate moiety. Peroxide migration involves elimination of an uncharged phosphate moiety with ring opening of the deoxyribose. Of course, radical oxidation of DNA bases can also occur. DNA peroxidation and cleavage leads to activation of PARP that activates
|
PO |
PO |
|
|
|
|
|
|
|
O |
O |
|
|
B |
|
|
|
B |
|
|
H |
.C |
|
HO. |
|
radical |
DNA cleavage |
PO |
cleavage |
|
|
|
PO |
|
|
|
|
–OP |
|
|
|
|
|
|
|
O2 |
|
PO |
|
|
|
O.
O
PO
PO
HOO
PO
Figure 10.1
O |
|
O+ |
B |
OP |
|
|
|
B |
|
|
|
|
|
|
HO– |
|
|
O |
DNA cleavage |
|
|
|
|
Criegee |
|
|
|
Rearrangement |
|
|
|
|
|
PO |
|
H. |
|
|
|
O |
PO |
|
|
|
|
|
|
B |
|
O |
|
|
|
|
B |
|
O |
|
DNA cleavage |
migration |
|
|
|
|
|
|
|
|
O |
|
|
|
P |
HO |
–OP |
DNA peroxidation and cleavage by oxygen radicals. B is any DNA base. PO is phosphate in the DNA structure. DNA cleavage occurs through three mechanisms: cleavage of the deoxyribose radical, Criegee rearrangement and peroxide migration.
DNA, Nuclear Cell Signaling and Neurodegeneration |
181 |
DNA repair enzymes.69 PARP uses NAD as an energy source and as a substrate to poly(ADP-ribosylate) itself and several nuclear enzymes. This poly(ADP-ribosylation) alters the activities of many enzymes.
Neuronal DNA is much more than just an archive of genetic information. It is very actively involved in producing proteins for neurotransmitter synthesis, release, reuptake and other neuronal functions. Damage to neuronal DNA very extensively a ects the ability of neurons to function normally, including maintaining neurotransmitters.70
PARP may ADP-ribosylate various transcription factors that regulate gene transcription.71,72 PARP is a component of positive cofactor 1 activity that regulates class II gene transcription.70 When DNA is damaged, PARP is activated, which inhibits class II gene transcription regulated by RNA polymerase II.73 PARP is also important in the action of p53, the tumor suppressor protein. Both PARP and p53 bind to DNA breaks. PARP can form complexes with p53 that may alter the activity of p53.74 Interestingly, p53 is involved in the inhibition of RNA polymerase III dependent gene transcription.75
It is important to recognize that the PARP referred to above is PARP-1. There are several enzymes with PARP activity.76 It is not known if the other enzymes, PARP-2, PARP-3, tankyrase and V-PARP, can fill in for PARP-1 when PARP inhibitors are used. Clearly, in PARP-1 knockout mice, the other PARP enzymes are still functional and can protect DNA and synthesize poly(ADP-ribose). It is also not known if PARP inhibitors are specific for PARP-1 or can inhibit all forms of PARP.
The energetic consequences of DNA damage and PARP activation are
enormous (Figure 10.2). PARP activation rapidly depletes NAD and ATP levels in the cell, leaving the cell depleted of energy sources.69,77 NADPH
depletion also occurs.77 Glutathione oxidizes. Glycolysis, the pentose phosphate pathway and mitochondrial energetics are a ected.
Recent research indicates that the secondary brain injury associated with stroke is induced by inflammatory processes. Nicotinamide can decrease the recruitment of neutrophils to potential sites of inflammation by inhibiting PARP in neutrophils and other cells.78 In fact, nicotinamide has been recommended for the treatment of arthritic patients since the 1940s. It was found in pilot trails that nicotinamide improved joint mobility and decreased the need for anti-inflammatory medication in arthritic cases.79 In the process of inflammation, the genes for intercellular adhesion molecule 1 and collagenase in neutrophils are activated. Neutrophils are recruited to sites of inflammation. Nitric oxide synthase is activated and oxygen radicals, hydrogen peroxide and nitric oxide are released. These reactive species can damage cellular DNA in the area of inflammation, resulting in apoptosis, necrosis and more serious inflammation.80 PARP has a number of functions in inflammation due to its ability to regulate gene expression.80 Inhibition by nicotinamide of PARP leads to decreased expression of these genes and decreases the extent and severity of inflammation. Nicotinamide also decreases the induction of iNOS, thereby decreasing damage to the blood-brain barrier.81

182 Chapter 10
|
|
|
|
|
|
|
|
|
|
|
|
|
Oxidative |
|
|
|
|
|
|
|||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
Insult |
|
|
|
|
|
|
|
|
|
||||
|
Loss of Mitochondrial function |
|
O2 |
|
|
|
|
|
|
|
|
|
||||||||||||||
|
|
|
|
|
|
|
|
|
|
|
||||||||||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|||||||||||||||
|
|
Lipid Peroxidation |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|||||||||
|
|
|
|
O2.- |
|
|
|
|
|
|
|
|
|
|||||||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
||||
Protein oxidation |
|
|
|
|
|
|
|
|
SOD |
|
|
|
|
|
|
|
|
|
||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
||
DNA PARP DNA |
|
|
|
|
|
|
|
|
|
|
GSH peroxidase |
|||||||||||||||
|
|
|
|
. |
Fe+2 |
|||||||||||||||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
H2O2 |
|
|
|
|
|
|
|
|
H2O |
||||
repair |
|
|
|
damage |
|
|
|
|
|
|
|
|
|
|||||||||||||
|
|
|
HO |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|||||||||
NAM |
|
NAD |
|
|
|
|
|
|
|
|
|
|
GSH |
|
GSSG |
|
|
|||||||||
|
|
|
|
|
|
NAD depletion |
GSSG Reductase |
|
|
|||||||||||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
NADP NADPH |
|||||||||
|
|
|
|
|
|
|
|
|
|
|
|
NAD kinase |
|
|
|
|
|
|
|
|
|
|||||
|
|
|
|
|
ATP |
NAD |
|
|
|
ATP |
PPP |
|
|
|||||||||||||
|
Na-AD |
|
|
Glycolysis |
G-6-P |
|
R-5-P |
|
|
|||||||||||||||||
|
|
|
|
|
||||||||||||||||||||||
|
|
|
2 |
|
|
|
|
2 |
|
|
ATP |
|
|
|
||||||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|||||||||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|||||||||
ATP |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
fructose |
|
|
|||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
Na-MN |
|
NMN |
|
|
|
|
|
Glucose |
|
|
|
||||||||||||||
ATP |
|
1 |
|
|
|
|
1 |
|
|
ATP |
|
|
|
|
|
|
NAD G- |
3-P |
|
|
||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
||||||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
||||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
NADH |
|
|
|
|
||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|||
|
|
|
|
|
|
|
|
NAM |
|
|
|
|
|
|
|
|
|
|||||||||
|
|
Na |
|
|
|
|
|
|
|
|
|
|
|
|
|
|||||||||||
|
|
|
|
|
|
|
NAD NADH |
|
|
2ATP |
||||||||||||||||
|
|
|
|
|
|
|
|
|
||||||||||||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
Lactate |
|
|
|
|
Pyruvate |
|||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|||||||||
Figure 10.2 The energetic consequences of DNA damage caused by oxidative stress. PARP is poly(ADP-ribose) polymerase. NAM is nicotinamide. NMN is nicotinamide mononucleotide. Na is niacin. Na-MN is niacin mononucleotide. Na-AD is niacin adenine dinucleotide. 1 is nicotinamide phosphoribosyl transferase, which requires ATP. 2 is NMN adenyl transferase, which requires ATP. PPP is the pentose phosphate pathway. G-6-P is glucose-6-phosphate. R-5-P is ribose-5-phosphate. G-3-P is 3-phosphoglycerate.
DNA damage leads to cell death. A large amount of DNA fragmentation causes rapid necrosis, with cell swelling and rupture, nuclear swelling and rupture and cytoplasmic vacuole formation. A smaller amount of DNA fragmentation causes apoptosis with cell shrinkage, nuclear condensation and fragmentation, large vacuole formation in the cytoplasm and cell fragmentation forming apoptotic bodies.82 Necrosis is a rapid process that does not require energy. Apoptosis is a delayed process that requires ATP. The center of a brain infarction is typically made up of necrotic cell debris.83 The limit area surrounding the core contains necrotic, apoptotic and normal cells.83
