
биохимия атеросклероза
.pdfxii Contributors
Bronwyn E. Brown, Free Radical Group, The Heart Research Institute, 114 Pyrmont Bridge Road, Camperdown, Sydney, NSW 2050, Australia
Jim W. Burgess, Liponex, Inc., 1740 Woodroffe Avenue, Ottawa, Ontario, Canada, K2G 3R8
Kanta Chechi, Department of Biochemistry, Memorial University of Newfoundland, St. John’s, Newfoundland, Canada
Sukhinder Kaur Cheema, Department of Biochemistry, Memorial University of Newfoundland, St. John’s, Newfoundland, Canada Email: skaur@mun.ca
Paul K.M. Cheung, Division of Stroke and Vascular Disease, St. Boniface General Hospital Research Centre, Department of Physiology, University of Manitoba, Winnipeg, Manitoba, Canada
Patrick C. Choy, Centre for Research and Treatment of Atherosclerosis, St. Boniface Hospital Research Centre, University of Manitoba, Winnipeg, Manitoba, Canada
Christophe M. Chretien, Liponex, Inc., 1740 Woodroffe Avenue, Ottawa, Ontario, Canada, K2G 3R8
Stephen M. Colgan, Department of Pathology and Molecular Medicine, McMaster University and the Henderson Research Centre, Hamilton, Ontario, Canada
John P. Cooke, Division of Cardiovascular Medicine, Stanford University School of Medicine, 300 Pasteur Drive, Stanford, California, USA Email: john.cooke@stanford.edu
Michael J. Davies, Free Radical Group, The Heart Research Institute, 114 Pyrmont Bridge Road, Camperdown, Sydney, NSW 2050, Australia Email: m.davies@hri.org.au
Naranjan S. Dhalla, Institute of Cardiovascular Sciences, St. Boniface General Hospital Research Centre and Department of Physiology, Faculty of Medicine, University of Manitoba, Winnipeg, Canada
Parambir Dhami, Institute of Cardiovascular Sciences, St. Boniface General Hospital Research Centre and Department of Physiology, Faculty of Medicine, University of Manitoba, Winnipeg, Canada
Victor A. Drover, Department of Pharmacological Sciences, State University of New York, Stony Brook, New York, USA
Pratibha Dubey, Department of Biochemistry, Memorial University of Newfoundland, St. John’s, Newfoundland, Canada
Contributors xiii
Gordon A. Francis, CIHR Group on Molecular and Cell Biology of Lipids, Departments of Medicine and Biochemistry, University of Alberta, Edmonton, Alberta, Canada
Email: gordon.francis@ualberta.ca
Manohar Garg, School of Biomedical Sciences, Faculty of Health, University of Newcastle, Callaghan, NSW 2308, Australia
Email: Manohar.Garg@newcastle.edu.au
Dean Gilham, Departments of Pediatrics and Cell Biology, University of Alberta, Edmonton, Alberta, Canada
Vicki Gill, Discipline of Medicine, Faculty of Medicine, Health Sciences Centre, Memorial University of Newfoundland, St. John’s, Newfoundland, Canada
Grant M Hatch, Department of Pharmacology and Therapeutics, Faculty of Medicine, Centre for Research and Treatment of Atherosclerosis,
St. Boniface Hospital Research Centre, University of Manitoba, Winnipeg, Manitoba, Canada
Email: hatchgm@Ms.UManitoba.CA
Gene R. Herzberg, Department of Biochemistry, Memorial University of Newfoundland, St. John’s, Newfoundland, Canada
Email: geneherz@mun.ca
Donald W. Jacobsen, Department of Cell Biology, Lerner Research Institute Department of Cardiovascular Medicine, Division of Medicine, Cleveland Clinic Foundation and Department of Molecular Medicine, Cleveland Clinic Lerner College of Medicine, Case Western Reserve University, Cleveland, Ohio USA
Richard Lehner, Departments of Pediatrics and Cell Biology, University of Alberta, Edmonton, Alberta, Canada
Email: richard.lehner@ualberta.ca
Lesley MacDonald-Wicks, School of Health Sciences, Faculty of Health, University of Newcastle, Callaghan, NSW 2308, Australia
Nandini Nair, Division of Cardiovascular Medicine, Stanford University School of Medicine, 300 Pasteur Drive, Stanford, California, USA
Kathryn E. Naus, Department of Pathology and Laboratory Medicine, University of British Columbia, Vancouver, British Columbia, Canada
Dominic S. Ng, Department of Medicine, St. Michael’s Hospital, West Annex 2-015, 38 Shuter Street, Toronto, Ontario, Canada
Email: ngd@smh.toronto.on.ca
xiv Contributors
Sereyrath Ngeth, CIHR Group on Molecular and Cell Biology of Lipids, Departments of Medicine and Biochemistry, University of Alberta, Edmonton, Alberta, Canada
Grant N. Pierce, Division of Stroke and Vascular Disease, St. Boniface General Hospital Research Centre, Department of Physiology, University of Manitoba, Winnipeg, Manitoba, Canada
Email: gpierce@sbrc.ca
Karmin O, Department of Animal Science, Faculty of Agricultural and Food Sciences, Department of Physiology, Faculty of Medicine, Centre for Research and Treatment of Atherosclerosis, National Centre for Agri-Food Research in Medicine, St. Boniface Hospital Research Centre, University of Manitoba, Winnipeg, Manitoba, Canada
Tanya A. Ramsamy, Departments of Pathology and Laboratory Medicine & Cellular and Molecular Medicine, University of Ottawa Heart Institute, Ottawa, Ontario, Canada
Imran Rashid, Free Radical Group, The Heart Research Institute, 114 Pyrmont Bridge Road, Camperdown, Sydney, NSW 2050, Australia
Manfredi Rizzo, Department of Clinical Medicine and Emerging Diseases, University of Palermo, Italy
Email: rizzomv@yahoo.it
Daisy Sahoo, Department of Pharmacological Sciences, State University of New York, Stony Brook, New York, USA
Email: daisysahoo@hotmail.com
Harjot K. Saini, Institute of Cardiovascular Sciences, St. Boniface General Hospital Research Centre and Department of Physiology, Faculty of Medicine, University of Manitoba, Winnipeg, Canada
P.K. Shah, Division of Cardiology and Atherosclerosis Research Center, Burns and Allen Research Institute and Department of Medicine, Cedars Sinai Medical Center and UCLA School of Medicine, Los Angeles, California, USA
Behrooz Sharifi, Division of Cardiology and Atherosclerosis Research Center, Burns and Allen Research Institute and Department of Medicine, Cedars Sinai Medical Center and UCLA School of Medicine, Los Angeles, California, USA
Email: sharifi@cshs.org
Garry X. Shen, Departments of Internal Medicine and of Physiology, University of Manitoba, Winnipeg, Manitoba, Canada
Email: gshen@ms.umanitoba.ca
Philip A. Sinclair, Liponex, Inc., 1740 Woodroffe Avenue, Ottawa, Ontario, Canada, K2G 3R8
Contributors xv
Yaw L. Siow, Department of Physiology, Faculty of Medicine, Centre for Research and Treatment of Atherosclerosis, National Centre for Agri-Food Research in Medicine, St. Boniface Hospital Research Centre, University of Manitoba, Winnipeg, Manitoba, Canada
Daniel L. Sparks, Liponex, Inc., 1740 Woodroffe Avenue, Ottawa, Ontario, Canada, K2G 3R8 and The Lipoprotein and Atherosclerosis Research Group, University of Ottawa, 40 Ruskin Street, Ottawa, Ontario, Canada Email: DSparks@ottawaheart.ca
Bernardo Trigatti, Department of Biochemistry and Biomedical Sciences, McMaster University, Hamilton, Ontario, Canada
Email: trigatt@mcmaster.ca
Hannah Valantine, Division of Cardiovascular Medicine, Stanford University School of Medicine, 300 Pasteur Drive, Stanford, California, USA
David M. van Reyk, Department of Health Sciences, University of Technology Sydney, PO Box 123, Broadway, NSW, 2007, Australia.
Sudesh Vasdev, Discipline of Medicine, Faculty of Medicine, Health Sciences Centre, Memorial University of Newfoundland, St. John’s, Newfoundland, Canada
Email: svasdev@mun.ca
Cheryl L.Wellington, Department of Pathology and Laboratory Medicine, University of British Columbia, Vancouver, British Columbia, Canada Email: cheryl@cmmt.ubc.ca
Geoff H. Werstuck, Departments of Biochemistry and Biomedical Sciences, Medicine, and the Henderson Research Centre, McMaster University, Hamilton, Ontario, Canada
Email: gwerstuck@thrombosis.hhscr.org
Stewart C. Whitman, Departments of Pathology and Laboratory Medicine & Cellular and Molecular Medicine, University of Ottawa Heart Institute, Ottawa, Ontario, Canada
Email: swhitman@ottawaheart.ca
Enoka P. Wijekoon, Department of Biochemistry, Memorial University of Newfoundland, St. John’s, Newfoundland, Canada
Yan-Jun Xu, Institute of Cardiovascular Sciences, St. Boniface General Hospital Research Centre and Department of Physiology, Faculty of Medicine, University of Manitoba, Winnipeg, Canada
Section I
Hyperlipidaemia and Atherosclerosis
Biochemistry of Atherosclerosis edited by S.K. Cheema, Springer, New York, 2006
1
Reverse Cholesterol Transport
JIM W. BURGESS, PHILIP A. SINCLAIR, CHRISTOPHE M. CHRETIEN, JONATHON BOUCHER, AND DANIEL L. SPARKS
Abstract
Atherosclerosis, the accumulation of cholesterol in the arteries resulting in heart attacks and strokes, is the leading cause of death in the USA and most other industrialized countries in the world. Plasma levels of high-density lipoprotein (HDL) cholesterol are invariably found to be inversely associated with the risk of atherosclerosis. This protective effect has classically been ascribed to HDL-mediated reverse cholesterol transport (RCT). In this process, nascent HDL in the circulation removes unesterified (free) cholesterol from peripheral cells, such as macrophages, through the transfer of cholesterol across the cell membrane by the ATP-binding cassette (ABCA1) transporter protein. The mature, spherical HDL particle, which contains these cholesterol esters in its core, then transfers the cholesterol to the liver through receptor-mediated processes. HDL cholesterol that is taken up by the liver is then excreted in the form of bile acids and cholesterol, completing the process of RCT. Because HDL-mediated RCT reduces serum cholesterol levels and is associated with a reduction in the risk of cardiovascular disease, increasing HDL is now being realized as a promising therapeutic end point.
Keywords: ABCA1; apolipoproteins; bile acids; cholesterol ester transfer protein; high-density lipoprotein; lipase; scavenger receptor BI
Abbreviations: ABC, ATP-binding cassette proteins; ACAT, acyl coenzyme A-cholesterol acyltransferase; apoA-I, apolipoprotein A-I; apoB, apolipoprotein B; AP1, transcription factor AP1; CE, cholesterol ester; CETP, cholesterol ester transfer protein; CYP7A1, cytochrome P450, family 7, subfamily A, polypeptide 1; EL, endothelial lipase; FXR, farnesoid X receptor; HDL, high-density lipoprotein; HNF- 4, hepatocyte nuclear factor-4; HL, hepatic lipase; HMGCoA, hydroxymethylglutaryl coenzyme A; JAK2, Janus kinase 2; LCAT, lecithin cholesterol acyltransferase; LXR, liver X receptor; LDL, low-density lipoprotein; LPL, lipoprotein lipase; LRP, LDLreceptor related protein; nCEH, neutral cholesterol ester hydrolase; PPAR, peroxisome proliferator activated receptor; RCT, reverse cholesterol transport; RXR, retinoid X receptor; SR-BI, scavenger receptor class B type I; Sp1, transcription factor Sp1; SREBP, sterol regulatory element binding protein; TG, triglyceride; StAR, steroidogenic acute regulatory protein; VLDL, very low-density lipoprotein
3

4 Jim W. Burgess et al.
Liver
Pre-β-HDL |
Lipases |
HDL |
|
||
apoA-I |
LCAT |
SR-BI |
|
FC CE |
|
FC |
|
|
CE
ABCA1 TG CETP CE
FC + Bile acids
LDL-R
Macrophage LDL
Feces
Intestine
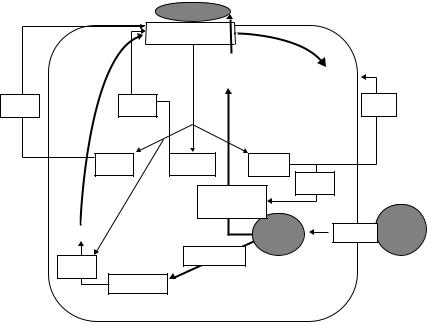
FIGURE 1.1. Schematic of the reverse cholesterol transport pathway. Cholesterol efflux to nascent HDL is regulated by ABCA1. Within the HDL pool, LCAT esterifies FC to CE and then CETP transfers CE from HDL to the LDL pool. FC and CE are taken up by the liver and recycled into the lipoprotein pool or secreted into the bile. ABCA1, ATP-binding cassette protein A1; apoA-I, apolipoprotein A-I; CE, cholesterol ester; CETP, cholesterol ester transfer protein; FC, free cholesterol; HDL, high-density lipoprotein; LCAT, lecithin cholesterol acyltransferase; LDL-R, low-density lipoprotein receptor; SR-BI, scavenger receptor class BI; TG, triglyceride.
Cholesterol that is synthesized in extrahepatic tissues or acquired from lipoproteins returns to the liver for excretion in a process called reverse cholesterol transport (RCT). RCT is a multistep process involving (i) cholesterol efflux from peripheral cells, (ii) maturation of the pre-β-high-density lipoprotein (pre-β-HDL) particle to spherical HDL, (iii) transfer of HDL-associated cholesterol to the liver, (iv) hepatic recycling or excretion of excess cholesterol, and (v) regeneration of the cholesterol acceptor pre-β-HDL. These individual steps are shown in Fig. 1.1 and will be reviewed separately in this chapter.
Cholesterol Efflux
The first step in RCT involves the transfer of cholesterol from cell membranes to acceptor lipoprotein particles in the extravascular fluids. Two mechanisms control this transfer of cholesterol. In the aqueous diffusion model, cholesterol molecules spontaneously disassociate from cell membranes, traverse the intervening aqueous space by diffusion, and then incorporate into acceptor particles. This process is driven by the gradient of free
Chapter 1. Reverse Cholesterol Transport |
5 |
cholesterol (FC) between cells and the aqueous medium and does not require energy or interaction of the cholesterol acceptor with a cell surface receptor [1]. The second mechanism requires interaction between lipid-poor HDL and cell surface ABCA1. ABCA1, an ATP-binding cassette transporter mutated in Tangier disease [2], promotes cellular phospholipid and cholesterol efflux by loading free apolipoprotein A-I (apoA-I) with these lipids. This process involves binding of apoA-I to the cell surface and the energy-dependent transfer of phospholipid and cholesterol by ABCA1 to apoA-I [3]. Crosslinking studies have suggested a very close association of apoA-I and ABCA1 at the cell surface [4, 5], however, there is also evidence that the mechanisms by which apoA-I associates with the plasma membrane and ABCA1 vary depending on cell type. Macrophages demonstrate a requirement for the C-terminal domain of apoA-I for optimal binding and efflux [6]. This may result from a binding site for the C-terminal domain of apoA-I on the macrophage extracellular matrix [7]. In contrast, apoA-I binding and efflux to wild-type apoA-I and several deletion mutants including those lacking residues 187–243 of the C-terminal tail [6] is equally efficient in human fibroblasts. Coincidentally, fibroblasts lack the extracellular matrix binding site [7]. By analogy with multidrug resistance proteins, also members of the ATP-binding cassette transporter family, it has been suggested that ABCA1 forms a channel within the plasma membrane through which phospholipids and cholesterol are transferred to lipid-poor particles [8].
In addition to stimulating lipid efflux from cells, the interaction of apoA-I and cell surface ABCA1 also turns on signal transduction pathways and their downstream effectors. A schematic model illustrating these processes is shown in Fig. 1.2. In fibroblasts, HDL or apoA-I has been shown to activate members of the Rho family of G-proteins [9] and downstream processes including adenylate cyclase and phospholipases including phosphatidylinosi- tol-specific phospholipase C and phosphatidylcholine-specific phospholipase D [10, 11]. Downstream protein kinases including protein kinase A (PKA) [12], protein kinase C (PKC) [13, 14], and Janus kinase 2 (JAK2) [15] are also activated. ApoA-I binding to human fibroblasts and consequent PKC activation is reported to mobilize intracellular cholesterol for efflux from an acyl coenzyme A-cholesterol acyltransferase (ACAT) accessible compartment [16]. Direct modulation of ABCA1 activity has been reported as a result of its phosphorylation. PKC directly phosphorylates ABCA1, which is reported to protect the protein against proteolytic degradation [17]. PKA activation by analogs of cAMP increase ABCA1 phosphorylation and cellular cholesterol efflux [18, 19]. Conversely, apoA-I-mediated release of cellular lipids is sensitive to inhibitors of PKA and impaired when PKA phosphorylation sites in ABCA1 are mutated [19]. Naturally occurring mutations of ABCA1 from Tangiers patients also demonstrated severely reduced apoA-I-mediated cAMP production, ABCA1 phosphorylation, apoA-I binding, and lipid efflux [18]. The protein-tyrosine kinase, JAK2, is required for optimum ABCA1-dependent lipid loading of apolipoproteins and appears to modulate
6 Jim W. Burgess et al.
|
Lipid-poor apoA-I |
|
|
|
|
|
|
|
ABCA1 |
|
|
|
|
|
|
Lipid efflux |
|
|
|
|
|
|
FC |
|
Degradation |
|
|
+ ve |
+ ve |
|
|
|
− Ve |
|
|
PKA |
JAK2 |
PKC |
|
|
|
|
|
Cholesterol |
+ ve |
|
|
|
|
|
|
|
|
||
|
cAMP |
mobilization |
|
|
|
|
|
Synthesis |
|
CE |
|
SR-AI |
apoB |
|
+ ve |
oxysterols |
|
|
|
|
|
|
Lipid droplet |
Modified LDL |
|||
|
LXR/RXR |
|
|
|
||
|
|
|
|
|
|
|
FIGURE 1.2. Regulation of the cholesterol efflux pathway. Binding of lipid-poor apoA-I to cell surface ABCA1 stimulates the efflux of FC and phospholipids and the activation of several protein kinases including PKA, PKC, and JAK2. PKA-mediated phosphorylation of ABCA1 optimizes the interaction between apoA-I and ABCA1 and increases the cholesterol translocase activity of ABCA1. Activation of JAK2 also enhances ABCA1 translocase activity. PKC-mediated phosphorylation of ABCA1 reduces degradation of ABCA1 and increases FC levels by mobilizing intracellular stores of cholesterol. The levels of ABCA1 are tightly controlled in cells. Synthesis is increased by metabolic products of lipid degradation that are ligands for the nuclear LXR/RXR receptor. Activated LXR/RXR increases transcription of the ABCA1 gene. ApoB, apolipoprotein B; JAK2, Janus kinase 2; LXR, liver X receptor; PKC, protein kinase C; PKA, protein kinase A; SR-AI, scavenger receptor class AI.
the apolipoprotein interactions with the ABCA1 transporter [15]. However, the lack of detectable phosphotyrosines in ABCA1 implies that the target of JAK2 is probably not ABCA1 [15].
The level of ABCA1 in cells is highly regulated. ApoA-I binding increases ABCA1 protein levels in mouse primary hepatocytes and peritoneal macrophages without affecting ABCA1 mRNA levels. This indicates that the rates of catabolism exert some control over ABCA1 levels in cells [20]. At the gene level the ABCA1 promoter contains binding motifs for several transcription factors, including the sterol regulatory element binding protein
Chapter 1. Reverse Cholesterol Transport |
7 |
(SREBP), the peroxisome proliferator activated receptor-α (PPAR-α), the liver X receptor (LXR), retinoid X receptor (RXR), and transcription factors Sp1 and AP1 binding sites [21]. These factors allow regulation through many diverse metabolites, including cAMP [5], sterols [22], cis-retinoic acid [23], PPAR agonists [24], and interferon-γ [25]. PPAR-α is a nuclear receptor activated by fatty acid derivatives and hypolipidemic drugs of the fibrate class. PPAR-α is expressed in monocytes, macrophages, and foam cells, suggesting a role for this receptor in macrophage lipid homeostasis, foam cell formation, and the consequent development of atherosclerosis [26].
Maturation of Nascent Pre-β-HDL
The transport of cholesterol from peripheral tissues to the liver by HDL (Fig. 1.1) is considered to be stimulated by the plasma enzymes lecithin cholesterol acyltransferase (LCAT) and cholesterol ester transfer protein (CETP). LCAT is synthesized in the liver and associates predominantly with HDL in the bloodstream. LCAT is activated by apoA-I [27] and converts HDL-associated cholesterol to cholesterol ester (CE). Nascent pre-β-HDL particles eventually become mature spherical particles after accumulation of sufficient CE. Spherical HDL is then acted on by CETP, which facilitates the transport of CE to the storage lipoproteins, low-density lipoprotein (LDL) and very low-density lipoprotein (VLDL), in exchange for TG [28]. LCAT and CETP therefore regulate the plasma storage of cholesterol and may contribute to RCT by channeling cholesterol from extrahepatic tissues to apolipoproteins B (apoB) that are subsequently catabolized in the liver [29]. Conversely, there is also data to suggest that inhibition of cholesterol storage pathways may stimulate RCT. Injection of phosphatidylinositol into the bloodstream of rabbits has been shown to inhibit LCAT activity and to directly stimulate RCT [30].
LCAT activity does not appear to be controlled at the gene level but is more affected by the composition and electrostatic properties of the plasma lipoproteins. In humans and some animal models on atherogenic diets, treatment with cholesterol-reducing drugs does not have a significant impact on LCAT expression [31]. In contrast, the human CETP promoter is under the control of several regulatory elements including the orphan nuclear receptor ARP1 [32], the CCAAT/enhancer-binding protein [33], trans-retinoic acid [34], the SREBP [35] and the liver receptor homolog-1 [36]. This allows several metabolic factors to modulate CETP levels including dietary cholesterol [37], fatty acids [38], and corticosteroids [39]. Numerous studies have reported that CETP modifies HDL cholesterol (HDL-C) and apoA-I levels, which are decreased by overexpression of CETP [40]. Human genetic deficiency of CETP results in significantly increased plasma HDL levels [41]. This has led to a considerable interest in CETP inhibitors as drugs to elevate HDL-C levels.
