
Genomics: The Science and Technology Behind the Human Genome Project. |
Charles R. Cantor, Cassandra L. Smith |
|
Copyright © 1999 John Wiley & Sons, Inc. |
|
ISBNs: 0-471-59908-5 (Hardback); 0-471-22056-6 (Electronic) |
4 Polymerase Chain Reaction
and Other Methods for In Vitro
DNA Amplification
WHY AMPLIFY DNA?
The importance of DNA signal amplification for sensitive detection of DNA through hybridization was discussed in the previous chapter. Beyond mere sensitivity, there are two
basic reasons why direct amplification |
of |
DNA is |
a |
vital part |
of |
DNA |
analysis. First, |
||
DNA amplification provides a route |
to |
an |
essentially |
limitless supply |
of |
material. When |
|||
amplification is used this way, as |
a |
bulk |
preparative |
procedure, |
the |
major requirement is |
|||
that the sample be uniformly amplified so that it is not altered, distorted, or mutated by the process of amplification. Only if these constraints can be met, can we think of amplification as a true immortalization of the DNA.
The second rationale behind DNA amplification is that selective amplification of a region of a genome, chromosome, or sample provides a relatively easy way to purify that segment from the bulk. Indeed, if the amplification is sufficient in magnitude, the DNA product becomes such an overwhelming component of the amplification mixture that the starting material is reduced to a trivial contaminant for most applications.
Amplification can be |
carried out in vivo by growing living cells (Box 1.2) or |
in |
vitro |
|
by using enzymes. There are |
several overwhelming advantages of in vitro amplification. |
|
|
|
Any possible toxic effects |
of a DNA target on the host cell are eliminated. There |
is |
no |
|
need to purify the amplified material away from the host genome or the vector used for |
|
|||
cloning. Base analogues |
can |
be used that would frequently be unacceptable to a living |
||
cell system. Samples can |
be |
manipulated by automated methods that are far easier to im- |
|
|
plement in vitro than in vivo. The major limitation of existing in vitro amplification meth-
ods until recently is that |
they |
were restricted to relatively short stretches of DNA, typi- |
||
cally |
less than 5 |
kb. |
New |
long polymerase chain reaction (PCR) procedures have |
extended the range of in |
vitro amplification up to about 20 kb. For longer targets than |
|||
this, |
in vivo cloning methods must still be used. |
|||
BASIC PRINCIPLES |
OF |
THE |
POLYMERASE CHAIN REACTION (PCR) |
|
What makes PCR a tool of immense power and flexibility is the requirement of DNA polymerases for pre-existing DNA primers. Thus DNA polymerases cannot start DNA
chains de novo; a primer can be used to determine where, along a DNA template, the synthesis of the complement of that stand begins. It is this primer requirement that allows the selective amplification of any DNA region by using appropriate, specific DNA primers.
98

|
|
|
BASIC PRINCIPLES OF THE POLYMERASE CHAIN REACTION (PCR) |
99 |
|||||||||||||||||
Once started, a DNA polymerase like |
|
|
|
|
E. coli |
|
DNA |
polymerase I |
(pol I) will proceed in |
|
|||||||||||
the 3 - to 5 |
-direction until it has |
copied all the |
way |
to |
the |
5 |
|
|
|
|
|
|
-end of the template. The |
||||||||
simplest amplification scheme for in vivo DNA amplification is successive |
cycles |
of |
|
|
|||||||||||||||||
priming, chain extension, and product denaturation, shown schematically in Figure 4.1. If |
|
|
|||||||||||||||||||
these steps are carried out efficiently, the result is a linear increase in the amount of prod- |
|
||||||||||||||||||||
uct strand with increasing cycles of amplification. This scheme, called |
|
|
|
|
|
|
|
linear |
amplifica- |
||||||||||||
tion, |
is very useful in preparing DNA samples for DNA sequencing. Here chain terminat- |
|
|
||||||||||||||||||
ing analogues of dpppN’s are added |
in trace amounts to produce |
a |
distribution |
of |
|
||||||||||||||||
products with different chain lengths (see Chapter 10). When linear amplification is used |
|
||||||||||||||||||||
in this context, it is called |
|
cycle sequencing. |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|||||
A step up in complexity from linear amplification |
is |
the |
use |
of |
two |
|
antiparallel |
|
|||||||||||||
primers, as shown in Figure 4.2 |
|
|
a . These primers define |
the region to be amplified, since |
|
||||||||||||||||
after the first few cycles of DNA synthesis, the relative amount of longer DNA sequences |
|
|
|||||||||||||||||||
that contain the region spanned by the primer becomes insignificant. The target DNA is |
|
|
|||||||||||||||||||
denatured, and two antiparallel primers are added. These must be in sufficient excess over |
|
|
|||||||||||||||||||
target that renaturation of the original |
duplex is improbable, and essentially |
all |
products |
|
|||||||||||||||||
are primers annealed to single-stranded templates. The |
|
first |
cycle |
of |
DNA |
synthesis |
|
|
|||||||||||||
copies both of the original template |
strands. Hence it doubles the |
number |
of |
targets |
|
||||||||||||||||
present in the starting reaction mixture. Each successive round of |
DNA |
denaturation, |
|
||||||||||||||||||
primer binding, and chain extension will, in principle, produce a further doubling of the |
|
|
|||||||||||||||||||
number of target molecules. Hence the amount of amplified product grows exponentially, |
|
|
|
||||||||||||||||||
as 2 n , where |
n |
is the |
number of |
amplification cycles. This is the basic |
design |
of a |
typical |
|
|||||||||||||
PCR procedure. Note that only the DNA sequence flanked by the two primers is |
ampli- |
|
|
||||||||||||||||||
fied (Fig. 4.2). |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
||
Early PCR protocols employed ordinary DNA polymerases such as the Klenow frag- |
|
|
|||||||||||||||||||
ment of |
E. |
coli |
DNA |
polymerase |
I |
(a |
truncated |
version |
of |
the |
natural |
protein |
with its |
|
|||||||
5 -exonuclease |
activity removed). The difficulty |
with this |
approach |
is |
that |
these |
enzymes |
|
|
||||||||||||
Figure 4.1 Linear amplification of DNA by cycles of repeated in vitro synthesis and melting.

Figure 4.2 |
Exponential amplification of DNA in vitro by the use of two antiparallel primers. |
(a) |
Polymerase chain |
reaction (PCR) with successive cycles of DNA synthesis and melting. |
(B) Only |
the DNA flanked by the primers is amplified. |
|
|
100
|
|
|
BASIC PRINCIPLES OF THE POLYMERASE CHAIN REACTION (PCR) |
101 |
|||||||
are easily denatured by the high temperatures needed |
to separate the two strands |
of |
du- |
|
|
||||||
plex DNA. Unlike DNA, however, proteins like DNA polymerases, once denatured, are |
|
|
|
||||||||
generally reluctant to renature. Thus, with each cycle of amplification, it was necessary to |
|
|
|||||||||
add fresh polymerase for efficient performance. This problem was relieved when DNA |
|
|
|
||||||||
polymerases from thermophilic organisms became available. The most widely |
used |
of |
|
|
|
||||||
these enzymes in PCR is from |
Thermis acquaticus, |
the |
Taq |
polymerase. This enzyme has |
|||||||
an optimal temperature for polymerization in the range of 70 to 75°C. It extends DNA |
|
||||||||||
chains at a rate of about 2 kb per minute. Most important, it is fairly resistant to the con- |
|
||||||||||
tinual cycles of heating and cooling required for PCR. The half-life for thermal denatura- |
|
|
|||||||||
tion of |
Taq |
polymerase is 1.6 h at 95°C, but it is less than 6 minutes at 100°C. When very |
|
||||||||
high denaturation temperatures are needed, |
as in the PCR amplification of very |
G |
|
C |
|||||||
rich DNA, |
it is sometimes useful to employ |
even more thermal-stable polymerases such |
|
|
|
||||||
as enzymes isolated from organisms that |
grow in the superheated water (temperature |
|
|
||||||||
above 100°C) near geothermal vents. Such enzymes are called |
|
|
|
vent |
polymerases. |
||||||
Examples are the enzyme from |
Thermococcus litoralis |
|
|
with a half-life of 1.8 h at 100°C |
|||||||
and the enzyme from |
Pyrococcus furiosis |
|
with a half-life of 8 h at 100°C. However, these |
||||||||
enzymes are not as processive as |
|
Taq |
polymerase. This means that they tend to fall off |
|
|||||||
their template more readily, which makes it more difficult to amplify long templates. |
|
|
|
|
|||||||
With |
Taq |
|
polymerase typical PCR cycle parameters are |
|
|
|
|
|
|||
|
|
|
DNA denaturation |
|
92–96°C for 30 to 60 seconds |
|
|
||||
|
|
|
Primer annealing |
|
|
55–60°C for 30 seconds |
|
|
|
||
|
|
|
Chain extension |
|
|
72°C for 60 seconds |
|
|
|
||
The number of cycles used depends on the particular situation and sample concentrations, |
|
|
|
||||||||
but typical values when PCR is efficient are 25 to 30 cycles. Thus the whole process takes |
|
|
|||||||||
about an hour. Typical sample and reagent concentrations used are |
|
|
|
|
|
||||||
|
|
|
Target |
10 15 mol, or down to as little as 1 mol |
|
||||||
|
|
|
Primer |
2 |
10 11 mol |
|
|
|
|
|
|
|
|
|
dpppN’s |
2 |
10 8 mol of each |
|
|
|
|
|
|
These concentrations imply that the reaction will eventually saturate once the primer and |
|
|
|||||||
dpppN’s are depleted. |
|
|
|
|
|
|
|
|
|
There are a few peculiarities of the |
Taq |
polymerase that must be taken into |
considera- |
|
|||||
tion in |
designing PCR |
procedures |
or experimental protocols based |
on |
PCR. |
Taq |
poly- |
||
merase has no 3 |
-proofreading exonuclease activity. Thus it can, and does, misincorporate |
|
|
||||||
bases. We will say more |
about this |
later. |
|
Taq |
polymerase does have a 5 |
-exonuclease ac- |
|
||
tivity. Thus it will nick translate, which is sometimes undesirable. However, mutants exist |
|
|
|||||||
that remove this activity, and they can be used if necessary. The most serious problem |
|
|
|||||||
generally |
encountered is |
that |
Taq |
polymerase |
can |
add |
an extra nontemplate-coded A |
to |
|
the 3 -end of DNA chains, as shown in Figure 4.3 |
|
|
|
a . It can also, perhaps using a related |
|
||||
activity, make primer dimers, which may or may not contain additional uncoded residues |
|
|
|||||||
as shown in Figure 4.3 |
|
b. This is a serious problem because once such dimeric primers are |
|
|
|||||
created, they are efficient substrates for further amplification. These primer dimers are a |
|
|
|||||||
major source of artifacts in PCR. However, they are usually short and can be removed by |
|
|
|||||||
gel filtration or other sizing methods to prevent their interference with subsequent uses of |
|
|
|||||||
the PCR reaction product. |
|
|
|
|
|
|
|
||
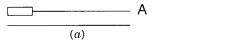
102 |
METHODS FOR IN VITRO DNA AMPLIFICATION |
||
Figure 4.3 |
Artifacts |
introduced |
by the use of |
Taq DNA polymerase. |
(a) |
Nontemplated terminal |
|
adenylation; |
(b)primer dimer formation. |
||
The efficiency of typical PCR reactions can be impressive. Let us |
consider a typical |
|
case where one chooses to amplify a short, specific region of human DNA. This sort of |
||
procedure would be useful, for example, in examining a particular region of the genome |
||
for differences among individuals, without having to clone the region from each person to |
||
be tested. To define this |
region, one needs to know the DNA sequence that flanks it, but |
|
one does not need know anything about the DNA sequence between primers. For unique, |
||
efficient priming, convergent primers about 20 bp long are used on each |
side of the re- |
|
gion. A typical amplification would start with 1 |
g of total human genomic DNA. Using |
|
Avogadro’s number, 6 |
1023, the genome size of 3 |
109 bp, and the 660 Da molecular |
weight of a single base pair, we can calculate that this sample contains
|
|
10 6 6 1023 |
|
3 105 |
copies |
|
|
3 109 6.6 102 |
|||
|
|
|
|
||
of the genome or 3 |
105 molecules of any single-copy genomic DNA fragments contain- |
||||
ing our specific target sequence. If PCR were 100% efficient, and 25 cycles of amplifica-
tion were carried out, we would expect |
to multiply the number of target |
molecules by |
||
225 6.4 107. Thus, after the amplification, the number of targets should be |
||||
6.4 |
107 3 105 2 |
1013 molecules |
||
If the product of the DNA amplification is 200 bp, it will weigh |
|
|||
200 bp |
|
(6.6 102 Da / bp) |
(2 1013) |
|
6 1023 g / Da |
||||
|
|
|||
4
g
If such efficient amplification could be achieved, the results would be truly impressive. This means that one would be sampling just a tiny fraction of the genome, and in a scant hour of amplification, the yield of this fraction would be such as to amount to 80% of the total DNA in the sample. Thus one would be able to purify any DNA sample.
In practice, the actual PCR |
efficiencies typically achieved are not perfect, but they are |
||||
remarkably good. We can define the |
efficiency, |
|
E |
, for n cycles of amplification by the ra- |
|
tio of product, |
P , to starting material, |
S , as |
|
||
|
|
|
P |
(1 E |
)n |
|
|
|
|
||
|
|
|
S |
|
|
Actual efficiencies turn out to be in the range of 0.6 to 0.9; this is impressive. Such high efficiencies immediately raise the notion of using PCR amplification as a quantitative tool
to measure not just the presence of a particular DNA sequence in a complex sample but to determine its amount. In practice, this can be done, but it is not always reliable; it usually requires coamplification with a standard sample, or competition with known amounts of a related sample.
NOISE IN PCR: CONTAMINATION |
103 |
NOISE IN PCR: CONTAMINATION
As in any high-gain amplification system, any fluctuations in conditions are rapidly mag-
nified, especially |
when |
they |
occur early in the reaction. There are many sources of noise |
||||||||
in PCR. One extremely common source is variations in the temperature at different posi- |
|||||||||||
tions in typical thermal cycling blocks. A typical apparatus for PCR uses arrays of tube |
|||||||||||
holders (or microtitre plate holders) that can be temperature controlled |
by |
heating |
ele- |
||||||||
ments and a cooling bath, by thermoelectric heating and cooling, by forced convection, or |
|
||||||||||
by switching among several pre-equilibrated water baths. A number of different funda- |
|||||||||||
mental designs for such thermal cyclers are now available, |
and |
the |
serious practitioner |
||||||||
would be well advised to look carefully at the actual temperature characteristics of the |
|||||||||||
particular |
apparatus used. |
For extremely |
finicky samples, |
such |
as |
those |
that |
need |
very |
||
G C rich primers, it may be necessary to use the same sample well each time to provide |
|||||||||||
reproducible PCR. It is for reasons like this that PCR, while it has revolutionized the han- |
|||||||||||
dling of DNA in research laboratories, has not yet found broad acceptance in clinical di- |
|||||||||||
agnostic laboratories despite its potential power. |
|
|
|
|
|
|
|||||
A major source of PCR noise appear to lie with characteristics of the samples them- |
|||||||||||
selves. The |
worst |
problem |
is undoubtedly sample contamination. If the same sample is |
|
|||||||
used repetitively in PCR assays, the most likely source of contamination is the PCR reac- |
|||||||||||
tion product from a previous assay. With PCR we are dealing with a system that amplifies |
|||||||||||
a DNA sequence by 10 |
7 fold. Thus, even if there is one part per million in carry over con- |
||||||||||
tamination; it will completely dominate the next round of PCR. A second major source of |
|||||||||||
contamination is DNA from organisms in dust or from DNA shed by the experimenter, in |
|
||||||||||
the form of dander, hair follicles, sweat, or saliva. This problem is obviously of greatest |
|||||||||||
significance when the samples to be amplified contain human sequences. |
|
|
|
|
|||||||
The basic cure for most contamination is to carry out PCR under typical biological |
|||||||||||
containment |
procedures |
and |
use good sterile technique including plugged pipetmen tips, |
||||||||
to prevent contamination by aerosols, and laminar flow hoods, to minimize the ability of |
|||||||||||
the investigator to inadvertently contaminate the sample. However, these approaches are |
|||||||||||
not always sufficient to deal with the contamination caused by previous PCR experiments |
|
||||||||||
on similar samples. This situation is extremely frustrating because it is not uncommon for |
|||||||||||
neophytes to have success with procedures that then progressively deteriorate as they gain |
|||||||||||
more experience, |
since |
the overall level |
and dispersal of |
contaminant in |
the |
laboratory |
|||||
keeps rising. There are general solutions to this problem, but they have not yet seen widespread adoption. One helpful procedure is to UV irradiate all of the components of a PCR reaction before adding the target. This will kill double-stranded DNA contaminants in the polymerase and other reagents. A more general solution is to exploit the properties of the enzyme, Uracil DNA glycosylase (DUG), which we described in Chapter 1. This enzyme
degrades DNA at each incorporated dU. Normally these |
incorporations are rare muta- |
|
genic events, and the lesion introduced into the DNA is rapidly repaired. |
||
DUG is used in PCR by carrying out the initial PCR amplification using dpppU instead of |
||
dpppT. The amplification product can be characterized in the |
normal |
way; the properties of |
DNA with T fully substituted by dU are not that abnormal except for |
a lower melting temper- |
|
ature and inability to be recognized by many restriction nucleases. Next, when a subsequent PCR reaction needs to be done, the sample, including the target, is treated with DUG prior to thermal cycling. This will destroy any carryover from the previous PCR because there will be
so many dU’s removed that the resulting DNA will be incapable of replication. If the second
PCR is also performed with dpppU, its subsequent carryover can also |
be prevented by a |
DUG treatment, and this procedure can be repeated |
ad libertum. |
104 METHODS FOR IN VITRO DNA AMPLIFICATION
PCR |
NOISE: |
MISPRIMING |
|
|
|
|
|
|
Typical PCR conditions with two convergent primers offer a number of |
possible |
unin- |
||||||
tended primed amplifications. These are illustrated in Figure 4.4. If the primers are not |
||||||||
chosen wisely, one of the two primers may be able to act alone to amplify DNA as shown |
||||||||
in Figure |
4.4 |
b. Alternatively, the two convergent primers may have more than one site in |
||||||
the target that allows amplification. There is no way to plan for these events, unless the |
||||||||
entire sequence of the sample is known. However, the chances of such accidental, unin- |
||||||||
tended, but perfect priming can be minimized by using long enough primers so that the |
||||||||
probability of such a coincidental match in DNA sequence is very small. |
|
|
||||||
|
A more serious and more common occurrence is mispriming by inexact pairing of the |
|||||||
primer with the template. Note that if the 3 |
|
-end |
of the primer is mispaired with the tem- |
|||||
plate, this is unlikely to lead to amplification, and thus there will be little harm. However, if |
||||||||
the 5 -end of the primer is mispaired, the impact is much more serious, as shown in Figure |
||||||||
4.4 |
c and |
|
d . If primer annealing is carried out under insufficiently |
stringent conditions, once |
||||
elongation is allowed to start, a 5 |
mispaired primer |
may |
still be able to lead to DNA syn- |
|||||
thesis. In the next round of PCR the incorrect elongated product from |
this synthesis will |
|||||||
serve as a template if it contains a sequence complementary to any of the primers in the so- |
||||||||
lution. However, when this synthesis extends past the original mispaired primer, the se- |
||||||||
quence that is made is now the precise complement of that primer. From this round on, no |
||||||||
stringency conditions will discriminate between the desired product and the misprinted arti- |
||||||||
fact. Thus more than one product will amplify efficiently in subsequent steps. The key is to |
||||||||
prevent the mispriming in the first place. If the |
misprimed |
sequence |
is |
nearly identical to |
||||
the |
primed |
sequence, the most obvious way to |
solve this |
problem is |
to change |
primers. |
||
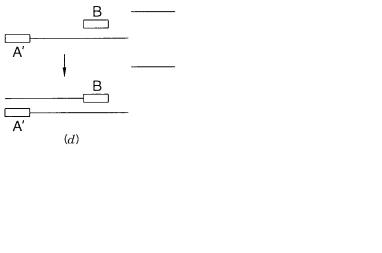
Figure 4.4 |
Effects of mispriming on |
PCR reaction products. |
(a) Desired product. |
(b)Product |
formed by an inverted repeat of one primer. |
(c) Product formed by nonstringent annealing. |
(d) After |
||
the second round of DNA synthesis, the product in |
(c) |
is now a perfect match to the primer for sub- |
|
|
sequent rounds of amplification. |
|
|
|
|

|
|
|
|
|
|
|
|
|
|
PCR |
NOISE: MISPRIMING |
105 |
|
Hopefully, another sequence near by on the desired template will not have its near mate |
|
|
|||||||||||
somewhere else in the sample. However, there are more general cures for some mispriming |
|
|
|
|
|||||||||
as shown below. |
|
|
|
|
|
|
|
|
|
|
|
||
|
In typical PCR reactions the denatured target and |
all |
of |
the |
other |
components |
are |
|
|
||||
mixed at room temperature, and then the cycles of amplification are allowed to start. This |
|
|
|
||||||||||
has the risk that polymerase extensions of the primers may start before |
the |
reaction |
is |
|
|
||||||||
heated to the optimal temperature for the elongation step. If this occurs, it enhances the |
|
|
|||||||||||
risk of mispriming, since room temperature is far from a stringent enough annealing tem- |
|
|
|
||||||||||
perature for most primers. This problem can easily be avoided by what is called |
|
|
|
hot |
start |
||||||||
PCR. |
Here the temperature is kept above the annealing temperature until all the compo- |
|
|
||||||||||
nents, including the denatured target, have been added. This avoids most of the misprim- |
|
|
|
||||||||||
ing during the first step; and it is always the first step in PCR that is most critical for the |
|
||||||||||||
subsequent production of undesired species. |
|
|
|
|
|
|
|
|
|
|
|||
|
A very powerful approach to favoring the amplification of desired products and elimi- |
|
|
|
|||||||||
nating the amplification of undesired products is nested PCR. This can be done whenever |
|
|
|
||||||||||
a sufficient length of known sequence is available at each end of the desired target. The |
|
|
|||||||||||
process is shown schematically in Figure 4.5. Two sets of primers are used; one is internal |
|
|
|
||||||||||
to the other. Amplification is allowed to proceed for half the desired rounds using the ex- |
|
|
|
||||||||||
ternal primers. Then the primers are switched to the internal primers, and amplification is |
|
|
|
||||||||||
allowed to continue for the remaining rounds. The only products |
that will be present at |
|
|
|
|||||||||
high |
concentration at the end of |
the reaction are those that |
can |
be |
amplified |
by |
both |
sets |
|
|
|
||
of primers. Any sequence that can inadvertently be amplified by |
one |
set |
is most |
unlikely |
|
|
|
||||||
to be a target for the second set, since, in general, there is no relationship or overlap of the |
|
|
|||||||||||
sequences used as the two sets of primers. With nested priming it is possible to carry out |
|
|
|||||||||||
many more than 30 rounds of amplification with relatively little background noise. Hence |
|
|
|
||||||||||
this procedure is to be especially recommended when very small |
samples |
are |
used and |
|
|
|
|||||||
large numbers of amplifications are needed to produce desired amounts of product. |
|
|
|
|
|
||||||||
|
Instead of full nesting, sometimes it is desirable or necessary |
to |
use |
a |
dual |
set |
of |
|
|||||
primers on one side of the target but only a single set on the other. This is |
called |
|
|
hemi- |
|||||||||
nesting, |
and it is still much safer and generally yields much cleaner products than no nest- |
|
|
||||||||||
ing at all. Particularly elegant versions of hemi-nesting have been demonstrated where the |
|
|
|
||||||||||
two |
nested primers can both be |
introduced at the start of |
the |
PCR |
at |
a temperature |
at |
|
|
||||
which only the external primer functions well. Halfway through the amplification cycles, |
|
|
|
||||||||||
the annealing temperature is shifted so that now the internal primer becomes by far the fa- |
|
|
|
||||||||||
vored one. |
|
|
|
|
|
|
|
|
|
|
|
||
Figure 4.5 The use of nested primers to increase the specificity of PCR amplification.
106 |
|
METHODS FOR IN VITRO DNA AMPLIFICATION |
|
|
|
|
|
|
|
|
|
|||||||
MISINCORPORATION |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
||
We mentioned earlier that |
|
Taq |
polymerase has no 3 |
|
editing exonuclease. Because of this |
|
|
|||||||||||
it has a relatively high rate of misincorporation when compared with many other DNA |
|
|
|
|
|
|
||||||||||||
polymerases. Thus the products of PCR reactions accumulate errors. The misincorpora- |
|
|
|
|
|
|
||||||||||||
tion |
rate |
of |
Taq |
polymerase has |
been |
estimated |
as 1.7 |
|
|
10 4 |
to 5 10 6 per nucleotide |
|||||||
per cycle. The error rate depends quite a bit on the reaction conditions, especially on the |
|
|
|
|
|
|||||||||||||
concentration of dpppN’s used, and on the sequence of the target. The impact of these mi- |
|
|
|
|
|
|||||||||||||
spairing rates on the product are straightforward to calculate. At any site in the target, the |
|
|
|
|
|
|||||||||||||
fraction of correct bases after n cycles of amplification will be |
|
|
|
|
|
|
|
|
|
|||||||||
|
|
|
|
|
|
X corr |
(1 X mis |
)n 1 nX |
mis |
|
|
|
|
|
||||
where |
|
X mis |
is the fraction of misincorporation rate at that site for a single cycle. For 30 cy- |
|
|
10 3 to |
||||||||||||
cles |
the |
fraction of misincorporated |
bases |
at any |
site |
will |
be |
30 |
|
|
X |
mis |
5.1 |
|||||
1.5 |
10 4 , using the numbers described above. Thus at any |
site the correct sequence is |
|
|
|
|||||||||||||
|
|
|
|
|||||||||||||||
still overwhelmingly predominant. |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
||||
However, if one asks, instead, for the amplification of a DNA of length, |
|
|
|
|
L , |
how |
many |
|||||||||||
incorrect bases will each DNA product molecule have after |
|
|
|
|
n |
steps, the number is |
|
LnX mis . |
||||||||||
With |
|
L 1000, |
this means that products |
will have |
from |
0.15 to 5 |
incorrect |
bases. How |
|
|
|
|
||||||
serious a problem is this? It depends on the use to which the DNA will be put. As a hy- |
|
|
|
|
|
|||||||||||||
bridization probe, these errors are likely to be invisible. If one sequences the DNA di- |
|
|
|
|
|
|||||||||||||
rectly, the errors will still be invisible (except for the rare case where a misincorporation |
|
|
|
|
|
|||||||||||||
occurred in the first round or two of the amplification and then was perpetuated). This is |
|
|
|
|
|
|
||||||||||||
because the errors are widely distributed at different sites on different molecules, and se- |
|
|
|
|
|
|||||||||||||
quencing sees only the average occupant of each site. However, if the PCR |
products |
are |
|
|
|
|
|
|||||||||||
cloned, the impact of misincorporation is much more serious. Now, since each clone is the |
|
|
|
|
|
|
||||||||||||
immortalization of a single DNA molecule, it will contain whatever particular errors that mol- |
|
|
|
|
|
|
||||||||||||
ecule had. In general, it is a hazardous idea to clone PCR products and then sequence them. |
|
|
|
|
|
|||||||||||||
The sequences will almost always have errors. Similarly PCR starting from single DNA mol- |
|
|
|
|
|
|
||||||||||||
ecules is fine for most analyses, but one cannot recommend it for sequencing because, once |
|
|
|
|
|
|
||||||||||||
again, the products are likely to show a significant level of misincorporation errors. |
|
|
|
|
|
|
|
|||||||||||
LONG PCR |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
||
A number of factors may limit the ability of conventional PCR to amplify long DNA tar- |
|
|
|
|
|
|
||||||||||||
gets. These include depurination of DNAs at the high temperatures used for denaturation, |
|
|
|
|
|
|
||||||||||||
inhibition of the DNA polymerase by stable intramolecular secondary structure in nomi- |
|
|
|
|
|
|
||||||||||||
nally single-stranded templates, insufficient time for strand untwisting during convention- |
|
|
|
|
|
|
||||||||||||
ally used denaturation protocols, as described in Chapter 3, and short templates. The first |
|
|
|
|
|
|||||||||||||
problem can be reduced by using increased pH’s to suppress purine protonation, a precur- |
|
|
|
|
|
|
||||||||||||
sor to depurination. The second problem can be helped somewhat by adding denaturants |
|
|
|
|
|
|
||||||||||||
like dimethyl sulfoxide (DMSO). The third problem can be alleviated by using longer de- |
|
|
|
|
|
|||||||||||||
naturation times. The last problem can be solved by |
preparing DNA |
in agarose (Chapter |
|
|
|
|
|
|
||||||||||
5). However, the most serious obstacle to the successful PCR amplification of long DNA |
|
|
|
|
|
|
||||||||||||
targets rests in the properties of the most commonly used DNA polymerase, the |
|
|
|
|
|
Taq |
poly- |
|||||||||||
merase. |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
SINGLE-SIDED |
PCR |
107 |
Taq |
polymerase |
has |
a |
significant |
rate of misincorporation as described above. |
|
|
|||||||||||
However, it lacks a 3 |
|
|
proofreading exonuclease activity. Once a base is misincorporated |
|
|
|||||||||||||
at the 3 -end, the chances that the |
extension |
will |
terminate |
at |
this |
point |
become |
|
|
|||||||||
markedly enhanced. This premature chain termination ultimately leads to totally ineffec- |
|
|
|
|||||||||||||||
tive PCR amplification above DNA sizes of 5 to 10 kb. To circumvent the problem of pre- |
|
|
|
|||||||||||||||
mature chain termination, Wayne Barnes and coworkers have added trace amounts of a |
|
|
|
|
||||||||||||||
second thermally stable DNA polymerase like |
|
|
|
|
|
|
Pfu, |
Vent, or Deep Vent that possesses a |
|
|||||||||
3 -exonuclease activity. This repairs any |
terminal mismatches |
left |
by |
|
|
|
|
Taq |
polymerase, |
|
||||||||
and then the latter can continue chain elongation. With such a two-enzyme procedure, |
|
|
|
|||||||||||||||
successful PCR amplification of DNA targets in the 20 kb to 40 kb range are now becom- |
|
|
|
|||||||||||||||
ing common. |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
INCORPORATING |
EXTRA FUNCTIONALITIES |
|
|
|
|
|
|
|
|
|
|
|
|
|
||||
PCR offers a simple and convenient way to enhance or embroider the properties of DNA |
|
|
|
|
||||||||||||||
molecules. As shown in Figure 4.6, primers can be pre-labeled with radioisotopes, biotin, |
|
|
|
|||||||||||||||
or fluorescent dyes. Thus the ends of the amplified targets can be derivatized as an intrin- |
|
|
|
|||||||||||||||
sic part of the PCR reaction. This is extremely |
convenient for many |
applications. Since |
|
|
|
|||||||||||||
two primers are chosen, two different labels or tags can be used. One frequent and every |
|
|
|
|||||||||||||||
effective strategy is to put a capture tag on one primer—like a biotin—and a detection tag |
|
|
|
|||||||||||||||
on the other—like a fluorophore. After the amplification, the product is captured and ana- |
|
|
|
|||||||||||||||
lyzed. Only double-stranded material that is the result of amplification that incorporated |
|
|
|
|||||||||||||||
both of the primers should be visible. |
|
|
|
|
|
|
|
|
|
|
|
|
|
|||||
PCR can also be used to modify the DNA sequence at the ends of the target. For exam- |
|
|
|
|||||||||||||||
ple, as shown in Figure |
4.6 |
|
b, |
the |
primer can overhang |
the ends |
of the desired target. As |
|
|
|||||||||
successive amplification cycles are carried out, the DNA duplexes that accumulate will |
|
|
|
|||||||||||||||
contain the target sequence flanked by the additional segments of primer sequence. This |
|
|
|
|||||||||||||||
has a number of useful applications. It allows any restriction sites needed to be built into |
|
|
|
|||||||||||||||
the primer. Then, as shown in the figure, |
after the PCR reaction the product can be |
|
|
|
||||||||||||||
cleaved at these sites for subsequent ligation or cloning steps. |
|
|
|
|
|
|
|
|
|
|||||||||
Another use for overhanging primers |
arises |
in |
circumstances |
where |
the original |
|
|
|
||||||||||
amount of known sequence is too short or too imperfect to allow efficient amplification. |
|
|
|
|||||||||||||||
This problem arises, for example, when a primer is made to an imperfectly repeating se- |
|
|
|
|||||||||||||||
quence. The usual desire in such experiments is to amplify many different copies of the |
|
|
|
|||||||||||||||
repeat (e.g., to visualize human DNAs among |
a background of rodent DNA in a hybrid |
|
|
|
|
|||||||||||||
cell as illustrated in Chapter 14), but few of the repeats match the primer well enough to |
|
|
|
|||||||||||||||
really give good amplification. By having |
an overhanging primer, after the first few |
|
|
|
||||||||||||||
rounds of amplification, the complements to the primer sequence now contain the extra |
|
|
|
|||||||||||||||
overhang (Fig. |
4.6 |
c ). The resulting template-primer complexes are much more stable and |
|
|
||||||||||||||
amplify much more effectively. |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|||
SINGLE-SIDED PCR |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
||
A major limitation in conventional PCR is that known DNA sequence is needed on both |
|
|
|
|||||||||||||||
sides of the desired target. It is frequently the case, as shown in Figure 4.7 |
|
|
a , that a known |
|
||||||||||||||
sequence |
is |
available |
only |
at |
one place |
within |
the |
desired |
target, |
or |
at |
one |
end of |
it. |
|
|
||
