
520 SEQUENCE-SPECIFIC MANIPULATION OF DNA
Figure 14.48 General construct needed for encoded combinatorial chemistry. Adapted from Brenner and Lerner (1992).
monomeric |
units in |
the oligomer will determine the complexity of the coding scheme |
needed in the DNA. A particularly nice trick, if small numbers of monomer units are in- |
||
volved, is |
to use a |
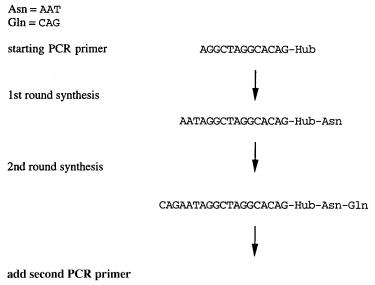
comma-less code. For example to specify the 20 amino acids, only a |
subset of 20 of the 64 possible triplet codons is needed. One can choose these, for example, so that if AAT and CAG represent two different amino acids, which can occur in ei-
ther order: AATCAG and CAGAAT, then ATC, TCA, AGA, and GAA are not assigned to
any amino acids. This makes the code resistant to frame shift errors and other ambiguities.
The chimeric DNA-oligomer compounds (Fig. 14.48) are screened for whatever activ-
ity is desired in the oligomer; then PCR is used just as in the select strategy in order to identify those components that have the desired affinity for a target. It is relatively easy to synthesize the full set of sequence identifiers and oligomers in a systematic way. This is
shown in Figure 14.49. The actual efficiency of such schemes needs to be tested experimentally. Undoubtedly new schemes and variations on existing schemes will proliferate.
However, |
the important |
feature of all of |
these approaches is that they |
illustrate |
the im- |
|
mense power that DNA analysis can bring to conventional chemistry. |
|
|
||||
OLIGONUCLEOTIDES |
AS |
DRUGS |
|
|
|
|
A large |
number of |
young biotechnology |
companies are betting their |
futures on |
the |
|
prospect that nucleic acids or nucleic acid analogs will function effectively as drugs. Most
of |
this effort is not based on conventional ideas about gene therapy, where a underactive |
or |
inactive defective gene might be supplemented by an active one, or an overactive or in- |
appropriately active gene might be substituted with a normal one. Such unconventional therapies are attractive, especially for many tissue-specific disorders, and such somatic

OLIGONUCLEOTIDES AS DRUGS |
521 |
Figure |
14.49 |
Split pool scheme for synthesizing a set of oligomeric compounds linked to their |
|||
specific oligonucleotide identifiers. Adapted from Brenner and Lerner (1992). |
|||||
gene therapy is already being tested in |
a few clinical trials. Here we are concerned with |
||||
the much more limited and conventional approach of attempting to use short pieces of nu- |
|||||
cleic acids or their analogues as drugs. |
|
|
|||
|
The intrinsic attractiveness of nucleic acids as drugs is their sequence specificity. One |
||||
can |
imagine that |
placed in the appropriate cell, an oligonucleotide could interfere with |
|||
RNA or DNA function either by binding |
directly to these species or by competing with |
||||
them for targets such as proteins. One approach is to simply use the sequence-specific |
|||||
binding |
affinity of |
the oligonucleotide to |
physically occlude a site or target. Here DNA is |
||
a potentially attractive target because |
it exists in very low copy numbers. An ideal anti- |
||||
gene scenario would be to design an oligonucleotide that would form a very stable triplex |
|||||
under physiological conditions with an unwanted promoter and so turn off the transcrip- |
|||||
tion |
of |
the gene |
controlled by this promoter. Alternatively, the oligonucleotide could be |
||
used |
as |
an affinity |
reagent to carry a photochemical or other covalent modifier to a target |
||
of interest. Again this seems potentially most effective with a DNA target. An extreme |
|||||
version of oligonucleotide therapy would be to use catalytic RNAs to find and destroy |
|||||
multiple target molecules. If this can be realized, it will be an extremely effective way to |
|||||
deal with infections by viruses with RNA genomes, or to attack other RNA targets. |
|||||
|
A number of |
obstacles must be overcome before successful oligonucleotide therapy |
|||
will |
be |
achieved. |
First, the materials |
must be delivered effectively to the correct target |
|
cells and in sufficient quantities to be therapeutically |
active. Side reactions with other |
||||
cells must be kept to a minimum. If cells lacked receptors for uptake of oligonucleotides, |
|||||
the |
problem would |
be to develop a targeting mechanism for |
the specific cells of interest. |
||

522 SEQUENCE-SPECIFIC MANIPULATION OF DNA
Unfortunately, at least some cells in the body, T lymphocytes, have a natural pathway for oligonucleotide uptake. How general this phenomenon is remains to be seen. The implication is that unless cells with intrinsic uptake pathways are the desired target, this uptake may have to be suppressed, possibly by competition with a harmless oligonucleotide and possibly by shielding the therapeutic compound in some way.
Once cellular uptake is achieved, the oligonucleotide must then be targeted successfully to the desired intracellular location. This will be the nucleus for a reagent directed against DNA, and it might be the nucleus or endoplasmic reticulum for reagents directed against RNA. Such targeting is not a simple matter. Most extracellular macromolecules taken up by cells are automatically targeted to the lysosome where they are destroyed. This natural pathway must be interfered with to successfully deliver a nucleic acid elsewhere. There is no doubt that one should be able to do this by exploiting the same sorts of processes that various viruses use to enter cells and infect the nucleus or the cytoplasm. However, many of these processes are not yet well understood, and we may have to learn
much more about them before successful oligonucleotide delivery mechanisms can be created.
Within the cell, or in intercellular fluids, a plethora of agents exist that can destroy or inactivate foreign nucleic acids. This is not surprising. Such agents must have evolved as antiviral defense mechanisms. To circumvent the action of these agents, oligonucleotide
drugs would either have to be introduced in large quantities or rendered resistant to a variety of nucleases and other enzymes of nucleic acid metabolism. For example, antisense messenger RNAs have been proposed as therapeutic agents to interfere with the translation of an unwanted message, arising perhaps from a virus or a tumor cell. These would be expected to act by binding to the normal mRNA and inactivating it for translation by occlusion or by a more active destructive process. The difficulty in vivo is the presence of
substantial amounts of RNA helicase activity. This is an enzyme that specifically recognizes double-stranded RNAs and unwinds the double helix. To be an effective drug under most circumstances, the backbone or bases of an antisense RNA will have to be altered so
that this molecule is no longer recognized by RNA helicases. |
|
|
|
Given the |
constrants mentioned above, the ideal oligonucleotide |
drug is |
probably |
likely to have an |
altered backbone to render it immune to normal nucleases |
and other |
en- |
zymes and to increase its binding affinity to natural nucleic acids. Compounds with uncharged backbones like PNAs seem particularly attractive in this regard. It may also be desirable to equip potential oligonucleotide-analog drugs with additional chemical func-
Figure 14.50 |
A potential |
oligonucleotide drug, designed by Claude Helene, that can bind to a |
DNA duplex and |
permanently inactivate |
it by photocrosslinking. For examples of results with this |
kind of approach, see Giovannangeli et al. (1992).
|
|
|
SOURCES AND ADDITIONAL |
READINGS |
|
523 |
|
tionalities in order to further enhance their binding and effectiveness at inactivating the |
|
|
|
||||
cellular target. An interesting example of such a potential drug is shown in Figure 14.50. |
|
|
|
||||
Designed by Claude Helene, this compound consists of a triplex-forming oligonucleotide |
|
|
|
||||
attached to a psoralen by a flexible chain. The psoralen enhances |
binding to DNA du- |
|
|
|
|||
plexes because it is an intercalator. More importantly, psoralen is a DNA photocrosslink- |
|
|
|
||||
ing, so near-UV irradiation, after formation of the triplex, results in irreversible crosslink- |
|
|
|||||
ing of the target DNA duplex. This procedure has |
been demonstrated to |
work effectively |
|
|
|
||
in cells. It may be a prototype of the sorts of materials that we will eventually see in ac- |
|
|
|
||||
tual therapeutic use. |
|
|
|
|
|
|
|
SOURCES |
AND ADDITIONAL READINGS |
|
|
|
|
|
|
Bailey, D. M. D., Carter, N. P., de Vos, D., Leversha, M. A., Perryman, M. T., and Ferguson-Smith, |
|
|
|||||
M. A. 1993. Coincidence painting: A rapid method for cloning region specific DNA sequences. |
|
|
|
||||
Nucleic Acids Research |
21: 5117 – 5123. |
|
|
|
|
||
Batzer, M. A., Alegria-Hartman, M., and Deininger, P. L. 1994. A consensus Alu repeat probe for |
|
|
|||||
physical mapping. |
Genetic Analysis: Techniques and Applications |
11: 34 – 38. |
|
||||
Brenner, S., and Lerner, R. A. 1992. Encoded combinatorial chemistry. |
Proceedings of the National |
|
|||||
Academy of Sciences USA |
89: 5381 – 5383. |
|
|
|
|
||
Brookes, A. J., and Porteous, D. J. 1992. Coincident sequence cloning: A new approach to genome |
|
|
|
||||
analysis. |
Trends in Biotechnology |
10: 40 – 44. |
|
|
|
|
|
DeRisi, J., Penland, P., Brown, P. O., Bittner, M. L., Meltzer, P. S., Ray, M., Chen, Y., Su, Y. A., and |
|
||||||
Trent, J. M. 1996. Use of a cDNA microarray to analyse gene expression patterns in human can- |
|
|
|
||||
cer. Nature Genetics |
14: 457 – 460. |
|
|
|
|
||
Eriksson, M., and Nielsen, P. E. 1996. PNA-nucleic acid complexes. Structure stability and dynam- |
|
|
|
||||
ics. Quarterly Review of Biophysics |
29: 369 – 394. |
|
|
|
|||
Feigon, J., Dieckmann, T., and Smith, F. W. 1996. Aptamer structures from A to Z. |
|
Chemistry |
and |
||||
Biology |
3: 611 – 617. |
|
|
|
|
|
|
Ferguson, J. A., Bowles, T. C., Adams, C. P., and Walt, D. R. 1996. A fiber-optic DNA biosensor |
|
|
|||||
microarray for the analysis of gene expression. |
|
Nature Biotechnology |
14: 1681 – 1684. |
|
|||
Ferrin, L. J., and Camerini-Otero, R. D. 1991. Selective cleavage of human DNA: RecA-assisted re- |
|
|
|||||
striction endonuclease (RARE) cleavage. |
Science |
254: 1494 – 1497. |
|
|
|||
Ferrin, L. J., and Camerini-Otero, R. D. 1994. Long-range mapping of gaps and telomeres with |
|
|
|||||
RecA-assisted restriction endonuclease (RARE) cleavage. |
Nature Genetics |
6: 379 – 383. |
|||||
Giovannangeli, C., Thuong, N. T., and Helene, C. 1992. Oligodeoxynucleotide-directed photo-in- |
|
|
|||||
duced cross-linking of HIV proviral DNA via triple-helix formation. |
|
Nucleic Acids Research |
20: |
||||
4275 – 4281. |
|
|
|
|
|
|
|
Gnirke, A., Iadonato, S. P., Kwok, P.-Y., and Olson, M. V. 1994. Physical calibration of yeast artifi- |
|
|
|||||
cial chromosome contig maps by recA-assisted restriction endonuclease (RARE) cleavage. |
|
|
|
||||
Genomics |
24: 199 – 210. |
|
|
|
|
|
|
Inque, S., Kiyama, R., and Oishi, M. 1996. Construction of highly extensive polymorphic DNA li- |
|
|
|||||
braries by in-gel competitive reassociation procedure. |
Genomics |
31: 271 – 276. |
|
||||
Irvine, D., Tuerk, C., and Gold, L. 1991. Selexion: Systematic evolution of ligands by exponential |
|
|
|||||
enrichment with integrated optimization by non-linear analysis. |
|
Journal of |
Molecular Biology |
|
|||
222: 739 – 761. |
|
|
|
|
|
||
Ito, T., Kito, K., Adati, N., Mitsui, Y., Hagiwara, H., and Sakaki, Y. 1994. Fluorescent differential |
|
|
|||||
display: Arbitrarily primed RT-PCR fingerprinting on automated DNA sequencer. |
|
FEBS Letters |
|||||
351: 231 – 236. |
|
|
|
|
|
||
524 |
SEQUENCE-SPECIFIC MANIPULATION OF DNA |
|
|
|
|
|
|||
Ito, T., Smith, C. L., and Cantor, C. R. 1992. Sequence-specific DNA purification by triplex affinity |
|
|
|||||||
capture. |
Proceedings of the National Academy of Sciences USA |
89: 495 – 498. |
|
||||||
Ito, T., Smith, C. L., and Cantor, C. R. 1992. Affinity capture electrophoresis for sequence-specific |
|
|
|||||||
DNA purification. |
|
Genetic Analysis: Techniques and Applications |
|
9: 96 – 99. |
|
|
|||
Ji, H., Smith, L. M., and Guilfoyle, R. A. 1994. Rapid isolation of cosmid insert DNA by triple- |
|
|
|||||||
helix-mediated affinity capture. |
Genetic Analysis: Techniques and Applications |
|
11: 43 – 47. |
||||||
Kato, K. 1996. RNA fingerprinting by molecular indexing. |
Nucleic Acids Research |
|
24: 394 – 395. |
||||||
Kandpal, R. P., Kandpal, G., and Weissman, S. M. 1994. Construction of libraries enriched for se- |
|
|
|||||||
quence repeats and jumping clones, and hybridization selection for region-specific markers. |
|
|
|
||||||
Proceedings of the National Academy of Sciences USA |
|
91: 88 – 92. |
|
|
|||||
Kool, E. T. 1996. Circular oligonucleotides: New concepts in oligonucleotide design. |
|
|
Annual |
||||||
Review of Biophysical and Biomolecular Structure |
25: 1 – 28. |
|
|
|
|||||
Liang, P., and Pardee, A. B. 1992. Differential display of eukaryotic messenger RNA by means of |
|
|
|
||||||
the polymerase chain reaction. |
Science |
257: 967 – 971. |
|
|
|
||||
Liang, P., Zhu, W., Zhang, X., Guo, Z., O’Connell, R. P. O. Averboukh, L., Wang, F., and Pardee, |
|
|
|||||||
A. B. 1994. Differential display using one-base anchored oligo-dT primers. |
|
|
Nucleic Acids |
||||||
Research |
22: 5763 – 5764. |
|
|
|
|
|
|||
Lisitsyn, N., Lisitsyn, N., and Wigler, M. 1993. Cloning the difference |
between two complex |
|
|
||||||
genomes. |
Science |
259: 946 – 951. |
|
|
|
|
|
||
Lockhart, D. J., Dong, H., Byrne, M. C., Follettie, M. T., Gallo, M. V., Chee, M. S., Mittmann, M., |
|
|
|||||||
Wang, C., Kobayashi, M. Horton, H., and Brown, E. 1996. Expression monitoring by hybridiza- |
|
|
|||||||
tion to high-density oligonucleotide arrays. |
Nature Biotechnology |
14: 1675 – 1680. |
|||||||
Mathieu-Daude, F., Cheng, R., Welsh, J., and McClelland, M. 1996. Screening of differentially am- |
|
|
|||||||
plified cDNA products from RNA arbitrarily primed PCR fingerprints using single strand con- |
|
|
|
||||||
formation polymorphism (SSCP) gels. |
|
Nucleic Acids Research |
24: 1504 – 1507. |
|
|||||
Moyzis, R. K., Torney, D. C., Meyne, J., Buckingham, J. M., Wu, J.-R., Burks, C., Sirotkin, K. M., |
|
|
|||||||
and Goad, W. B. 1989. The distribution of interspersed repetitive DNA sequences in the human |
|
|
|
||||||
genome. |
Genomics |
4: 273 – 289. |
|
|
|
|
|
||
Nallur, G. N., Prakash, K., and Weissman, S. M. 1996. Multiplex selection techniques (MuST): An |
|
|
|||||||
approach to clone transcription factor binding sites. |
|
Proceedings of |
the National Academy |
of |
|||||
Sciences USA |
93: 1184 – 1189. |
|
|
|
|
|
|||
Nielson, P. E., Egholm, M., Berg, R. H., and Buchardt, O. 1991. Sequence-selective recognition of |
|
|
|||||||
DNA by strand displacement with a thymine-substituted polyamide. |
|
Science |
254: 1497 – 1500. |
||||||
Perry-O’Keefe, H., Yai, X.-W., Coull, J. M., Fuchs, M., and Egholm, M. 1996. Peptide nucleic acid |
|
|
|||||||
pre-gel hybridization: An alternative to Southern hybridization. |
|
Proceedings of |
the |
National |
|||||
Academy of Sciences USA |
93: 14670 – 14675. |
|
|
|
|
||||
Pollock, R., and Treisman, R. 1990. A sensitive method for the determination of protein-DNA bind- |
|
|
|
||||||
ing specificities. |
|
Nucleic Acids Research |
18: 6197 – 6204. |
|
|
|
|||
Qureshi, S. J., Porteous, D. J., and Brookes, A. J. 1994. Alu-based vectorettes and splinkerettes |
|
|
|||||||
more efficient and comprehensive polymerase chain reaction amplification of human DNA from |
|
|
|
||||||
complex sources. |
|
Genetic Analysis: Techniques and Applications |
|
11: 95 – 101. |
|
||||
Rosenberg, M., Przybylska, M., and Straus, D. 1994. “RFLP subtraction.” A method for making li- |
|
|
|||||||
braries |
of polymorphic |
markers. |
Proceedings of the National |
Academy of Sciences USA |
|
91: |
|||
6113 – 6117. |
|
|
|
|
|
|
|
||
Schena, M. Shalon, D., Davis, R. W., and Brown, P. O. 1995. Quantitative monitoring of gene ex- |
|
|
|||||||
pression patterns with a complementary DNA microarray. |
|
Science |
270: 467 – 470. |
||||||
Sosnowski, R. G., Tu, E., Butler, W. F., O’Connell, J. P., and Heller, M. J. 1997. Rapid determina- |
|
|
|||||||
tion of |
single |
base |
mismatch mutations in DNA hybrids by direct electric |
field control. |
|
|
|
||
Proceedings of the National Academy of Sciences USA |
|
94: 1119 – 1123. |
|
|
|||||
|
|
|
|
SOURCES |
AND |
ADDITIONAL |
READINGS |
525 |
Straus, R., and Ausubel, F. M. 1990. Genomic subtraction for cloning DNA corresponding to dele- |
|
|
||||||
tion mutations. |
|
Proceedings of the National Academy of Sciences USA |
|
|
87: 1889 – 1893. |
|
||
Strobel, S. A., Doucette-Stamm, L. A., Riba, L., Housman, D. E., and Dervan, P. B. 1991. Site- |
|
|||||||
specific cleavage of human chromosome 4 mediated by triple-helix formation. |
|
|
Science |
254: |
||||
1639 – 1642. |
|
|
|
|
|
|
|
|
Takabatake, T. et al. 1992. The use of purine-rich oligonucleotides in triplex-mediated DNA isola- |
|
|
||||||
tion and generation of unidirectional deletions. |
Nucleic Acids Research |
20: 5853–5854. |
|
|||||
Unrau, P., and Deugau, K. V. 1994. Non-cloning amplifications of specific DNA fragments from |
|
|
||||||
whole genomic DNA digests using DNA “indexers.” |
|
Gene |
145: 163 – 169. |
|
||||
Velculescu, V. E., Zhang, L., Vogelstein, B., and Kinzler, K. W. 1995. Serial analysis of gene ex- |
|
|||||||
pression. |
Science |
270: 484 – 487. |
|
|
|
|
|
|
Veselkov, A. G., Demidov, V. V., Nielsen, P. E., and Frank-Kamenetskii, M. D. 1996. A new class of |
|
|||||||
genome rare cutters. |
|
Nucleic Acids Research |
24: 2483 – 2487. |
|
||||
Wan, J. S., Sharp, S. J., Poirier, G. M.-C., Wagaman, P. C., Chambers, J., Pyati, J., Hom, Y.-L., |
|
|||||||
Galindo, J. E., Huvar, A., Peterson, P. A., Jackson, M. R., and Erlander, M. G. 1996. Cloning |
|
|||||||
differentially expressed mRNAs. |
Nature Biotechnology |
|
14: 1685 – 1691. |
|
||||
Wittung, P., Kim, S. K., Buchart, O., Nielsen, P., and Norden, B. 1994. Interactions of DNA binding |
|
|||||||
ligands with PNA-DNA hybrids. |
Nucleic Acids Research |
|
22:5371 – 5377. |
|
||||
Wittung, P., Eriksson, M., Lyng, R., Nielsen, P., and Norden, B. 1995. Induced chirality in PNA- |
|
|||||||
PNA duplexes. |
|
Journal of the American Chemical Society |
|
|
117: 10167 – 10173. |
|
||
Yokota, H., and Oishi, M. 1990. Differential cloning of genomic DNA: Cloning of DNA with an al- |
|
|
||||||
tered primary structure by in-gel competitive reassociation. |
|
|
|
Proceedings of the National |
|
|||
Academy of Sciences USA |
87: 6398 – 6402. |
|
|
|
|
|||
