
Dale_Molecular Genetics of Bacteria 4th ed
.pdf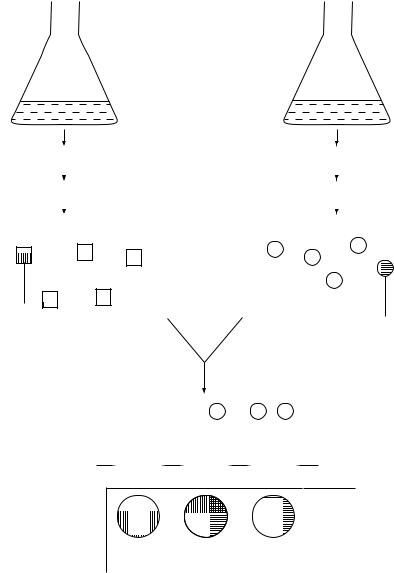
GENE MAPPING TO GENOMICS |
297 |
A B
|
mRNA |
|
mRNA |
|||||||||||||||||||||||||||||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
cDNA |
|
|
cDNA |
|||||||||||||||||||||||||||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
Label 1 |
|
Label 2 |
|||||||||||||||||||||||||||||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
Hybridize to microarray
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
3 |
|
|
|
|
|
1 |
|
|
|
|
2 |
|
|
|
|
|
|
|
|
|||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
||
|
|
|
Red |
|
|
|
Yellow |
|
|
Green |
||||||||||
Microarray (small portion)
Expression |
Expression |
Expression |
in A not B |
in both A |
in B not A |
|
and B |
|
Figure 10.12 Analysis of gene expression using a microarray. cDNA probes from cultures A and B are labelled with different fluorescent dyes, as described in Figure 10.8. If a gene is expressed more in culture A, label 1 will bind more strongly to that spot on the array, which will appear red. A gene expressed more strongly in culture B will show as green, while if there is no difference the spot will be shown as yellow
298 |
MOLECULAR GENETICS OF BACTERIA |
conditions. For this purpose, a cell extract would be fractionated by acrylamide gel electrophoresis (SDS PAGE), the proteins transferred to a filter and the specific proteins detected with antibodies. These antibodies are either labelled directly or detected by using a second labelled antibody which reacts with the first one. The most common label is an enzyme (such as peroxidase) that can be visualized using a chromogenic substrate. This procedure is known as Western blotting, by analogy with the Southern blotting technique described in Chapter 2, although the process is rather different as it usually involves electrophoretic transfer. However, Western blotting is only suitable for monitoring the expression of specific proteins. For global analysis of protein expression (the proteome), twodimensional gel electrophoresis may be employed.
Analysis of gene expression using proteomics
Whilst microarray technology allows the determination of the amount of all individual mRNAs in a cell, the scope of proteomics is to characterize the expression of every single protein in a cell at a specific time and under specific conditions. Bacterial cells may contain thousands of different proteins, so powerful methods are needed to separate them. The amino acid sequence of a protein determines both the mass and the overall charge of the molecule and these two characteristics can be used to separate the individual proteins of a cell. For example, many proteins have a similar mass and could not therefore be resolved by SDS-PAGE alone. However, most of those with similar mass have different compositions of acidic and basic amino acids which confer a different overall electrical charge to the protein. The overall charge of the protein depends on the pH. At high pH values they will be negatively charged, while at low pH the charge will be positive. In between is the isoelectric point (pI) at which the protein is neutral. The pI varies from one protein to another, depending on the amino acid composition.
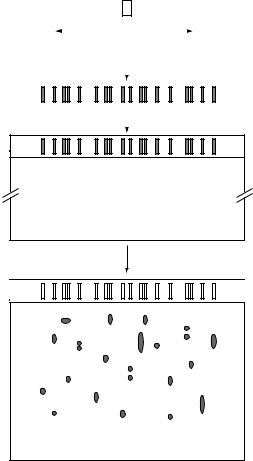
Two-dimensional polyacrylamide electrophoresis (2D-PAGE) separates proteins in two sequential steps: first by their charge and secondly by their mass (Figure 10.13). The first step involves isoelectric focusing (IEF) during which the proteins present in the cell extract are resolved on a gel in which a stable pH gradient is generated using chemicals called ampholytes. When an electric field is applied to the gel, each protein will migrate through the gradient until it reaches a point (the pI) at which the net charge of the protein is zero and it becomes stationary. Because they have different pI values, many of the proteins will travel to different positions on the IEF gel. However the resolution is not complete, as some proteins will have similar isoelectric points. Further resolution is achieved by applying the IEF gel to an SDS-PAGE gel, so as to separate the proteins on the basis of their mass alone. Using this technology as many as 1000 proteins can be resolved simultaneously.
|
GENE MAPPING TO GENOMICS |
|
299 |
|||||
|
|
|
Isoelectric focusing |
|
|
|||
|
|
|
|
|
|
|
|
|
+ |
Low |
|
|
|
|
High |
− |
|
pH |
|
|
|
|
pH |
|
||
|
|
|
|
|
|
|||
|
|
|
|
|
|
|
|
|
|
Negatively-charged Positively-charged |
|
|
|||||
|
proteins |
|
proteins |
|
|
|||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
SDS-PAGE gel
Separation by size
Figure 10.13 2D protein gel electrophoresis
After the gel has been run, a stain is applied to highlight the resolved proteins which appear as individual and separated spots. The location of a known protein in the final gel gives it a unique signature that is formed by its charge and mass and this can be used to identify it. Alternatively, the identity of an unknown protein can be determined by cutting the protein spot from the gel and subjecting it to micro-chemical analysis such as direct amino acid sequencing or mass spectrometry. These techniques will provide partial but unique amino acid sequences, which can then be linked back to the corresponding gene in the genome by reverse translation. Using 2D-PAGE it is now possible to study the total expression profile of nearly all the proteins in a bacterial cell under any chosen condition.
300 |
MOLECULAR GENETICS OF BACTERIA |
10. 4. 3 Systematic analysis of gene function
The genome sequence represents a catalogue of all the genes of a bacterium but many of the genes identified in these projects do not have a known function. Consequently, following the completed genome sequences for some bacteria, systematic programs to identify the function of all known reading frames have been initiated. The aim of these is to inactivate all the genes (where this is possible) of an individual species and by assessing the resulting phenotype, to determine the function of every individual protein. These ambitious and far-reaching projects are only now possible with the availability of complete bacterial genome sequences.
10.5 Conclusion
These final chapters can give no more than a brief overview of the range of techniques available and the extent to which molecular genetics has enhanced our understanding of the basic biology of bacteria (to say nothing of the revolution that has occurred with higher organisms). The expansion of the subject shows no signs of abating. The rapidity with which genome sequencing is advancing will by itself have a major impact on our understanding of all aspects of bacteriology. This includes not only the molecular biology of these organisms, but their physiology, biochemistry, pathogenicity, evolution – nothing escapes. However, in order to make the most of this information, the information gleaned from molecular studies has to be related back to the behaviour of the cell as a whole and its interaction with its environment. It is this synthesis of different fields that is the major challenge in the years ahead.

Appendix A
Further Reading
Comment
This is a highly selective list of the major texts that we find most useful – the absence of a book from this list does not necessarily mean that it has no merit.
In a rapidly moving field such as molecular genetics, information rapidly becomes out-of-date. It is not always wrong, but sometimes the language changes subtly or the emphasis alters or a tentative explanation becomes an accepted truth. So, a student relying too heavily on old books can convey a rather old-fashioned feel to the answers. The most recent edition should always be used and anything more than a few years old should not be referred to unless you are really sure of your ground. Wherever possible, this list should be supplemented with review articles and original literature. We have not included any such references as they date even more rapidly than do textbooks; however, if the reader wishes to search them out for themselves, Trends in Genetics/
Microbiology or Microbiological Reviews are recommended. Scientific American is also a valuable source of information. Some journals (such as Molecular Microbiology) also publish short review articles as well as research papers. For the more ambitious, reference to the Annual Reviews series (Biochemistry, Microbiology or Genetics) is suggested – but these articles do tend to be rather heavy going.
Symposium proceedings are often of limited use for student reading; the Society for General Microbiology symposia are a notable exception.
General texts and basic molecular biology
B.Lewin. Genes VII. (2000). Oxford University Press. First choice for further reference, but it may be found to be too big a jump from the level of this book. Wide ranging, but very useful for bacteria as well as eukaryotes.
C.R. Calladine and H. R. Drew. Understanding DNA: The Molecule and How It Works. 2nd edn (1997). Academic Press. (3rd edn due in 2004). An unusual book. Aimed partly at non-scientists, it starts in a deceptively simple manner, but progresses to dealing with many difficult concepts relating to the structure and function of DNA. Especially useful on supercoiling. Strongly recommended as an antidote to some of the rather superficial accounts of DNA structure in most molecular biology textbooks.
Molecular Genetics of Bacteria, 4th Edition by Jeremy Dale and Simon F. |
Park |
# 2004 John Wiley & Sons, Ltd ISBN 0 470 85084 1 (cased) ISBN 0 |
470 85085 X (pbk) |
302 |
FURTHER READING |
B.Alberts et al. Molecular Biology of the Cell, 4th edn (2002). Garland. Remarkable value for money, but difficult to carry around. Although relating primarily to eukaryotes, a valuable reference source for some of the molecular aspects.
H. Lodish et al. Molecular Cell Biology, 4th edn (1995). W. H. Freeman. (5th edn due in 2003–2004). Another text aimed primarily at eukaryotic cell biology, but containing enough information of general relevance, or specifically about bacteria, to be useful for reference on the broader molecular aspects.
More focus on bacteria
L.Snyder and W. Champness (2003). Molecular Genetics of Bacteria, 2nd edn. American Society for Microbiology. First choice for further reference on bacterial genetics.
S.Baumberg (ed.) (1999). Prokaryotic Gene Expression. A multi-author book, which makes for a very detailed account of selected topics rather than a fully integrated storyline. A big jump from the level of this book.
M. T. Madigan, J. M. Martinko and J. Parker (2000). Biology of Microorganisms (better known as ‘Brock’), 9th edn. Prentice Hall International. Included in case the reader needs to revise the biological background to some of the systems described.
Gene cloning
J.W. Dale and M. von Schantz (2002). From Genes to Genomes. John Wiley & Sons. Companion to this book, taking recombinant DNA technology much further and into the world of eukaryotes.
T.A. Brown (2001). Gene Cloning – An Introduction, 4th edn. Blackwell Science. An excellent introductory book.
S.B. Primrose, R. Twyman and R. W. Old (2001). Principles of Gene Manipulation, 6th edn. Blackwell Science. Much heavier going, but mandatory for the more advanced student.
B.R. Glick (2003). Molecular Biotechnology: Principles and Applications of Recombinant DNA, 3rd edn. American Society for Microbiology.
Websites which give more information on bacterial genome sequences and access to genomic data bases
http://www.sanger.ac.uk/ The website for the The WellcomeTrust Sanger Institute is one of the leading genomic centres in the world.
http://www.tigr.org/ The website for the Institute for Genomic Research (TIGR) which is involved in the comparative analysis of genomes and gene products from a wide variety of organisms including bacteria.
http://www.ncbi.nlm.nih.gov/ A website that contains public databases and software tools for analysing genome data.
FURTHER READING |
303 |
General genetics
Some suggestions for those who want to explore further the relationship between bacterial genetics and the broader field of genetics.
D. P. Snustad and M. J. Simmons (2000). Principles of Genetics, 2nd edn. John Wiley. W. S. Klug and M. R. Cummings (2000). Concepts of Genetics, 6th edn. Prentice Hall. L. H. Hartwell and others (2000). Genetics. McGraw-Hill.
P. J. Russell (2002). Genetics. Benjamin Cummings.
A.J. F. Griffiths, W. M. Gelbart, R. C. Lewontin and J. H. Miller (2002). Modern Genetic Analysis, 2nd edn. W. H. Freeman.
Special topics
R.W. Hendrix et al (1983). Lambda II. Although well over our usual time limit, this is still an invaluable source of reference about bacteriophage , but only for the more dedicated student at this level.
M. Wilson, R. McNab and B. Henderson (2002). Bacterial Disease Mechanisms. Cambridge University Press. Provides further information about many of the virulence mechanisms that provide important examples of the applications of bacterial genetics.
A special mention
W. Hayes (1968). The Genetics of Bacteria and their Viruses, 2nd edn. Blackwell Scientific Publications. The classic text. Recommended for those interested in the historical development of the subject – it still contains a surprising amount of useful material.

Appendix B
Abbreviations
Amino acid notations
|
Three-letter |
One letter |
|
notation |
notation |
Alanine |
Ala |
A |
Arginine |
Arg |
R |
Asparagine |
Asn |
N |
Aspartate |
Asp |
D |
Cysteine |
Cys |
C |
Glutamate |
Glu |
E |
Glutamine |
Gln |
Q |
Glycine |
Gly |
G |
Histidine |
His |
H |
Isoleucine |
Ile |
I |
Leucine |
Leu |
L |
Lysine |
Lys |
K |
Methionine |
Met |
M |
Phenylalanine |
Phe |
F |
Proline |
Pro |
P |
Serine |
Ser |
S |
Threonine |
Thr |
T |
Tryptophan |
Trp |
W |
Tyrosine |
Tyr |
Y |
Valine |
Val |
V |
Other abbreviations
5-BU |
5-Bromouracil |
A |
Adenine |
BAC |
Bacterial artificial chromosome |
Molecular Genetics of Bacteria, 4th Edition by Jeremy Dale and Simon F. |
Park |
# 2004 John Wiley & Sons, Ltd ISBN 0 470 85084 1 (cased) ISBN 0 |
470 85085 X (pbk) |
306 |
ABBREVIATIONS |
C |
Cytosine |
cAMP |
Cyclic AMP |
CAP |
Catabolite activator protein (see CRP) |
ccc |
Covalently closed circular (plasmid DNA) |
cDNA |
Complementary DNA |
CHEF |
Contour-clamped homogeneous electrophoretic field electrophoresis |
CRP |
Cyclic AMP receptor protein |
ddNTP |
20,30-Dideoxynucleoside triphosphate; any of ddATP, ddCTP, ddGTP or |
|
ddTTP (or a mixture of all four) |
DGGE |
Denaturing gradient gel electrophoresis |
dNTP |
Any one of dATP, dCTP, dGTP or dTTP (or a mixture of all four) |
DR |
Direct repeat |
ds |
Double-stranded (RNA or DNA) |
e.o.p. |
Efficiency of plating (of a bacteriophage) |
EMS |
Ethyl methane sulphonate |
EtBr |
Ethidium bromide |
FIGE |
Field inversion gel electrophoresis |
f-Met |
Formyl-methionine |
G |
Guanine |
GFP |
Green fluorescent protein |
GSP |
General secretory pathway |
Hfr |
High frequency of recombination |
HPK |
Histidine protein kinase |
IEF |
Isoelectric focusing |
IPTG |
Iso-propylthiogalactoside. |
IR |
Inverted repeat |
IS |
Insertion sequence |
IVET |
In vivo expression technology |
kb |
Kilobase (¼1000 bases) |
Mb |
Megabase (1 million bases) |
MNNG |
1-Methyl-3-nitro-1-nitroso-guanidine |
mRNA |
Messenger RNA |
NTP |
any one of ATP, GTP, CTP, or UTP (or a mixture of all four) |
oc |
Open circular (nicked form of plasmid DNA) |
ORF |
Open reading frame |
PAGE |
Polyacrylamide gel electrophoresis |
PCR |
Polymerase chain reaction |
PEG |
Polyethylene glycol |
PFGE |
Pulsed field gel electrophoresis |
pfu |
Plaque-forming units |
RF |
Replicative form |
pI |
Isoelectric point (pH at which a protein has no net charge) |
RFLP |
Restriction fragment length polymorphism |
RR |
Response regulator |
