
- •Introduction
- •Spotting phosphotyrosine
- •v-Src and other protein tyrosine kinases
- •Processes mediated through tyrosine phosphorylation
- •Tyrosine kinase-containing receptors
- •The ErbB receptor family and their ligands
- •Cross-linking of receptors causes activation
- •Assembly of receptor signalling complexes
- •Protein domains that bind phosphotyrosines and the assembly of signalling complexes
- •Branching of the signalling pathway
- •The Ras signalling pathway
- •From Ras to MAP kinase and the activation of transcription
- •Raf genes
- •Beyond ERK
- •Docking sites and a MAP kinase phosphorylation motif
- •Activation of protein kinases by ERKs 1 and 2
- •Activation of early response genes
- •Regulation of the cell cycle
- •Fine tuning the Ras-MAP kinase pathway: scaffold proteins
- •MAP kinase scaffold proteins discovered in yeast
- •KSR, a mammalian scaffold protein that regulates MAP kinase signalling
- •Other proteins that regulate MAP kinase pathways
- •Why are the signalling pathways so complicated?
- •Termination of the ERK response
- •Activation of PI 3-kinase
- •Direct phosphorylation of STAT transcription factors
- •A switch in receptor signalling: activation of ERK by 7TM receptors
- •Pathway switching mediated by receptor phosphorylation
- •Pathway switching by transactivation
- •Pathway switching, transactivation, and metastatic progression of colorectal cancer
- •References

Signalling Pathways Operated by Receptor Protein Tyrosine Kinases
phosphorylation assay, the V599E mutation enhances activity 700-fold, and this gives rise to a 2–5-fold increase in the activity of ERK1.109
The reason why only B-Raf, but not A- or C-Raf, is linked to tumour formation may be due to a difference in the N-terminal region. In B-Raf this carries an acidic sequence and phosphorylation is not needed in order to confer a negative charge. Fewer changes are needed to cause deregulation.
Beyond ERK
Docking sites and a MAP kinase phosphorylation motif
ERKs 1 and 2 phosphorylate substrates within the sequence Ser/Thr-Pro. Many proteins carry such a sequence, yet not all are ERK substrates, so there must be other determinants of specificity. Many kinases and phosphatases are regulated by interactions with their substrates at non-catalytic regions, docking motifs that help to avoid indiscriminate phosphorylations or dephosphorylations.
The idea that MAP kinases form stable links with their substrates first arose with studies of JNK2, which was isolated from cell lysates in tight association with its substrate c-Jun. The delta domain (D domain), characterized by a stretch of positively charged residues surrounded by a zone of hydrophobicity in c-Jun, is required for its stable association with JNK2.110,111 D domains are also involved in the interaction of ERKs1 and 2 with the kinases MEK1, RSK1, MSK, MNK1,
the dual specificity phosphatases MKP-3, MKP-3, MKP-7, and the transcription factor Elk-1.112 Some proteins, such as JunD, possess both DEF and
D domains.113 In others, a DEF motif is involved in the interaction of ERKs1 and 2 with transcription factors and also with the scaffold protein KSR1. The DEF motif seems to be reserved for substrates of the ERK family (not JNK or p38).
Finally, MAP kinases themselves possess conserved domains through which they communicate with their regulators or substrates (Figure 12.14). The common docking domain (CD), which is negatively charged, interacts with proteins possessing the positively charged D domain. Another is the ERK-docking or DE motif. Exchange of only two residues within the DE motif of ERK2 alters its binding specificity to that of p38, thereby switching the
attention of ERK2 to MK3.114 Such domain swaps cause a redirection of signals so that stress signals transmute into growth factor signals and vice versa.115
Activation of protein kinases by ERKs 1 and 2
Besides initiating transcription, ERKs1 and 2 also activate the MAP kinaseactivated protein kinases, MK (see Table 12.3). There are five subfamilies (Figure 12.15), all characterized by the presence of a CaMK-like kinase domain
ERKs 1 and 2 are widely coexpressed. They appear to phosphorylate similar substrates at
the same steps in the pathway, and they are similarly controlled. In accordance with others, we will henceforth refer to ERKs 1 and 2. However, it should be
noted that these kinases are not necessarily interchangeable.
337
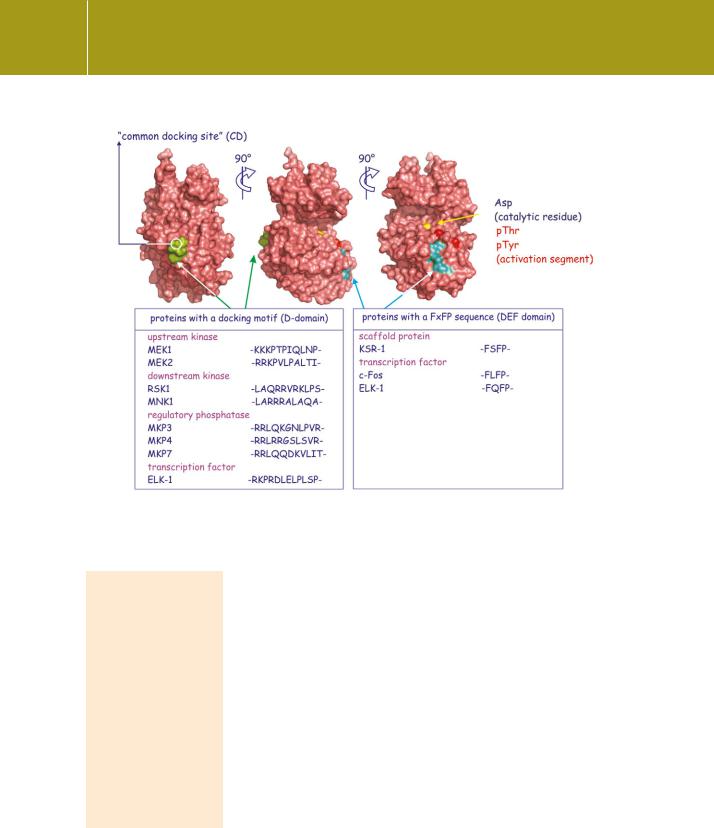
Signal Transduction
Fig 12.14 Molecular surface of ERK2 showing docking domains. ERK2 binds through its common docking site (CD, green) to proteins that possess a D domain. Proteins that possess a DEF domain bind at the other site on ERK (cyan), close to its activation segment (red spots). (2erk116) Adapted from Dimitri et al.117
RSK and S6K. RSK1 was first identified in extracts of Xenopus
laevis oocytes as a kinase that phosphorylates the ribosomal protein S6 (component of the 40S subunit).120 Later it was shown that S6 is actually a poor substrate for RSK, but a good substrate for the S6K family of protein kinases (S6Ks 1 and 2). The S6Ks are phosphorylated and activated by mTOR and PDK1 but not by the ERKs (see Chapter 18).
and all having similar activation segment sequences that are substrates of either ERKs1 and 2 or p38 kinases, or both. Two subfamilies, RSK1–4 and MSK1 and 2 carry a second catalytic domain in the N-terminal region which resembles that of PKC. Surprisingly, the activation segment of this domain is not a substrate of ERKs1 and 2 (nor of p38) so it is unclear how these kinases are activated. It is possible that phosphorylation by ERKs 1 and 2 prepares the MK target kinase for a second, activating phosphorylation by the membraneassociated PDK1 (see page 550).
RSK
The RSKs are present in the cytosol but move partially to the nucleus when activated by phosphorylation. They have specific substrates in both locations. In the cytosol, these are initiation factors of protein translation, components of the apoptosis machinery, the oestrogen receptor, and the guanine nucleotide exchange factor Sos. In the nucleus the RSKs phosphorylate ATF4, c-Fos, and SRF and generally enforce the ERK signal.119
338

Signalling Pathways Operated by Receptor Protein Tyrosine Kinases
Table 12.3 Substrates of the MAP kinases and MAP kinase-activated kinases (MK)
“(a) Substrates of MAP kinase”
|
ERK 1 & 2 |
p38 , , , |
JNK1, 2, 3 |
|
|
|
|
|
|
|
|
Membrane |
CD120a, Syk, calnexin |
|
|
|
|
|
|
|
|
|
|
Transcriptional |
SRC-1, Pax6, NF-AT, Elk- |
ATF1/2, MEF2a, |
c-jun, JunD, ATF-2, NF-Acc1, HSF-1, |
|
|
regulation |
1, MEF2, c-Fos, c-Myc, |
Sap-1, Elk-1, NF- |
STAT3, p53 |
|
|
(nucleus) |
STAT3, JunD, Ets-2 |
B, Ets-1, p53 |
|
|
|
|
|
|
|
|
|
Actin cytoskeleton |
Neurofilament, paxillin, |
Tau |
|
|
|
and focal |
vinexin |
|
|
|
|
adhesion |
|
|
|
|
|
|
|
|
|
|
|
Signalling |
MEK, Lin-1, RSK, MSK, |
PLA2, MNSK, MSK, |
Shc |
|
|
|
MNK, TSC2 |
MK2 |
|
|
|
|
|
|
|
||
(b) Substrates of MAP kinase-activated kinases (MK) |
|
|
|
||
|
|
|
|
|
|
|
RSK1–4 |
MSK1 & 2 |
MNK 1 &2 |
MK2 & 3 |
MK5 |
|
|
|
|
|
|
Transcription |
TIF-1, ER81, ER , Mi, |
Histone H3, |
ER |
CREB, SRF, |
|
|
CBP, CREB, c-Fos, c-Jun, |
HMG14, CREB, |
|
ER81,E47 |
|
|
Nur77, SRF, ATF-4 |
ATF1, p65, ER81, |
|
|
|
|
|
STAT3 |
|
|
|
|
|
|
|
|
|
RNA stability, |
|
4E-BP1 |
eIF-4E |
hnRNP, TTP, |
|
translation |
|
|
|
PABP1, HuR, |
|
|
|
|
|
eEFkinase |
|
|
|
|
|
|
|
Cell cycle |
P27kip, Myt1, Bub1 |
|
|
|
|
|
|
|
|
|
|
Cell survival |
Bad, LKB1, GSK3 , IkB , |
Bad |
|
|
|
|
C/EBP |
|
|
|
|
|
|
|
|
|
|
Signalling |
Sos, TSC2 |
PKB |
|
|
|
|
|
|
|
|
|
Miscellaneous |
|
|
|
glycogen |
|
|
|
|
|
synthase,14- |
|
|
|
|
|
3-3 , 5-LO |
|
Information from Roux and Blenis.118
MSK
MSKs is present mainly in the nucleus, where the substrates are CREB (see page 248), histone H3, HMGN1, and ATF1. It acts to modify DNA, making it more accessible and again, facilitates the response initiated by ERKs 1 and 2.118 Some of the substrates are listed in Table 12.3. For further details, see Roux and Blenis118 and Hauge and Frodin119.
339

Signal Transduction
Fig 12.15 Domain architecture of the MAP kinase-activated kinases, MK. These comprise the ribosomal S6 kinases (RSK), the mitogen and stress-activated kinases (MSK), the MAP kinase-interacting kinases (MNK), MAP kinase-activated protein kinases 2 and 3 (MK-2 and -3), and MAP kinase-activated protein kinase 5B (MK5B). Phosphorylations at the indicated sites are necessary for activation. ERK and p38 only phosphorylate the C-terminal activation segment. Members of families I and II possess a pair of non-identical kinase domains. Of these, the N-terminal domains phosphorylate substrate and are classified functionally as AGC kinases. Strangely enough, this domain is not phosphorylated by ERK. PDK1 is a potential candidate and phosphorylation of the C-terminal kinase by ERK may have a permissive role.
340

Signalling Pathways Operated by Receptor Protein Tyrosine Kinases
Fig 12.16 eIF-4E and the translation initiation complex. The initiation complex is formed around the eukaryotic initiation factor eIF-4E which is attached to the 5 -m7G cap of the mRNA. The initiation complex comprises tRNAMet (first amino acid), numerous other eukaryotic initiation factors, and the small ribosomal particle 40S. Phosphorylation of eIF-4E, by MNK1, facilitates complex formation and therefore accelerates protein translation.
MNK
The third subfamily of kinases activated by ERKs 1 and 2 is the MAP kinaseintegrating kinases, MNKs 1 and 2.121 MNK1 exerts an indirect stimulatory effect on protein synthesis through ERKs 1 and 2 and directly through phosphorylation of initiation factor-4E (eIF-4E). This is attached to the 5 -m7G cap of mRNA and acts as an anchor for the assembly of the initiation complex. It plays an important role in the binding of the small (40S) ribosomal subunit to the mRNA and in the subsequent search for the AUG start codon (Figure 12.16). MNK1 is associated with eIF-4G, a component of the initiation complex, and
341
