22.5 ● The Molecular Architecture of Photosynthetic Reaction Centers |
723 |
22.5 ● The Molecular Architecture of
Photosynthetic Reaction Centers
What molecular architecture couples the absorption of light energy to rapid electron-transfer events, in turn coupling these e transfers to proton translocations so that ATP synthesis is possible? Part of the answer to this question lies in the membrane-associated nature of the photosystems. Membrane proteins have been difficult to study due to their insolubility in the usual aqueous solvents employed in protein biochemistry. A major breakthrough occurred in 1984 when Johann Deisenhofer, Hartmut Michel, and Robert Huber reported the first X-ray crystallographic analysis of a membrane protein. To the great benefit of photosynthesis research, this protein was the reaction center from the photosynthetic purple bacterium Rhodopseudomonas viridis. This research earned these three scientists the 1984 Nobel Prize in chemistry.
Structure of the R. viridis Photosynthetic Reaction Center
Rhodopseudomonas viridis is a photosynthetic prokaryote with a single type of photosystem. The reaction center (145 kD) of the R. viridis photosystem is localized in the plasma membrane of these photosynthetic bacteria and is composed of four different polypeptides, designated L (273 amino acid residues), M (323 residues), H (258 residues), and cytochrome (333 amino acid residues). L and M each consist of five membrane-spanning -helical segments; H has one such helix, the majority of the protein forming a globular domain in the cytoplasm (Figure 22.16). The cytochrome subunit contains four heme groups; the N- terminal amino acid of this protein is cysteine. This cytochrome is anchored to the periplasmic face of the membrane via the hydrophobic chains of two fatty acid groups that are esterified to a glyceryl moiety joined via a thioether bond to the Cys (Figure 22.16). L and M each bear two bacteriochlorophyll molecules (the bacterial version of Chl) and one bacteriopheophytin. L also has a bound quinone molecule, QA. Together, L and M coordinate an Fe atom. The photochemically active species of the R. viridis reaction center, P870, is composed of two bacteriochlorophylls, one contributed by L and the other by M.
Photosynthetic Electron Transfer in the R. viridis Reaction Center
The prosthetic groups of the R. viridis reaction center (P870, BChl, BPheo, and the bound quinones) are fixed in a spatial relationship to one another that favors photosynthetic e transfer (Figure 22.16). Photoexcitation of P870 (creation of P870*) leads to e loss (P870 ) via electron transfer to the nearby bacteriochlorophyll (BChl).The e is then transferred via the L bacteriopheophytin (BPheo) to Q A, which is also an L prosthetic group. The corresponding site on M is occupied by a loosely bound quinone, Q B, and electron transfer from Q A to Q B takes place. An interesting aspect of the system is that no elec-
Cytochrome with
 4 heme groups
4 heme groups
hν
|
M |
L |
|
|
P870 |
|
|
<1ps |
|
BChl |
BChl |
|
BPheo |
20ps |
|
|
|
|
BPheo |
|
|
230ps |
|
|
QA |
|
QB |
Fe |
|
100µs |
|
|
H
Note: The cytochrome subunit is membraneassociated via a diacylglycerol moiety on its N-terminal Cys residue:
O
+ H 
 H3N C C
H3N C C
CH2
FIGURE 22.16 ● Model of the structure and activity of the R. viridis reaction center. Four polypeptides (designated cytochrome, M, L, and H) make up the reaction center, an integral membrane complex. The cytochrome maintains its association with the membrane via a diacylglyceryl group linked to its N-terminal Cys residue by a thioether bond. M and L both consist of five membrane-spanning -helices; H has a single membranespanning -helix. The prosthetic groups are spatially situated so that rapid e transfer from P870* to Q B is facilitated. Photoexcitation of P870 leads in less than 1 picosecond (psec) to reduction of the L-branch BChl only. P870 is re-reduced via an e provided through the heme groups of the cytochrome.
Q
2H+
QH2
1–2 H+
FIGURE 22.17
3–4 H+
Cytb/c1
724 Chapter 22 ● Photosynthesis
Bacterial
F1F0 ATP synthase
tron transfer occurs through M, even though it has components apparently symmetrical to and identical with the L e transfer pathway.
The reduced quinone formed at the Q B site is free to diffuse to a neighboring cytochrome b/cytochrome c1 membrane complex, where its oxidation is coupled to H translocation (and, hence, ultimately to ATP synthesis) (Figure 22.17). Cytochrome c2, a periplasmic protein, serves to cycle electrons back to P870 via the four hemes of the reaction center cytochrome subunit. A specific tyrosine residue of L (Try162) is situated between P870 and the closest cytochrome heme. This Tyr is the immediate e donor to P870 and completes the light-driven electron transfer cycle. The structure of the R. viridis reaction center (derived from X-ray crystallographic data) is modeled in Figure 22.18.
Eukaryotic Reaction Centers: The Molecular Architecture of PSII
PSII of higher plants and green algae contains more than 20 subunits and is considerably more complex than the R. viridis reaction center. Nevertheless, the R. viridis reaction center is a fairly good model for the core structure of PSII. P680 and its two equivalents of pheophytin (Pheo) are located on a pair of integral membrane proteins designated D1 (38 kD) and D2 (39.4 kD) (Figure 22.19). The tyrosine species D is Tyr161 in the D1 amino acid sequence. Complexed to D2 is a tightly bound plastoquinone molecule, Q A. Electrons flow from P680* to Pheo on D1 and thence to Q A on D2 and then on to a second plastoquinone situated in the Q B site on D1. Electron transfer from Q A and Q B is assisted by an iron atom located between them. Each plastoquinone (PQ) that enters the Q B site accepts two electrons derived from water and two H from the stroma before it is released into the membrane as the hydroquinone, PQH2. The stoichiometry of the overall reaction catalyzed by PSII is
3 H+ 
CytC2
Outside
Light
CytC2





 M
M 

 L
L
2e–
QB Fe QA
H
2 H+
Cytoplasm
● The R. viridis reaction center is coupled to the cytochrome b/c1 complex through the quinone pool (Q). Quinone molecules are photoreduced at the reaction center Q B site (2 e [2 h ] per Q reduced) and then diffuse to the cytochrome b/c1 complex, where they are reoxidized. Note that e flow from cytochrome b/c1 back to the reaction center occurs via the periplasmic protein cytochrome c2. Note also that 3 to 4 H are translocated into the periplasmic space for each Q molecule oxidized at cytochrome b/c1. The resultant proton-motive force drives ATP synthesis by the bacterial F1F0 ATP synthase.
(Adapted from Deisenhofer, J., and Michel, H., 1989. The photosynthetic reaction center from the purple bacterium Rhodopseudomonas viridis. Science 245:1463.)
22.5 ● The Molecular Architecture of Photosynthetic Reaction Centers |
725 |
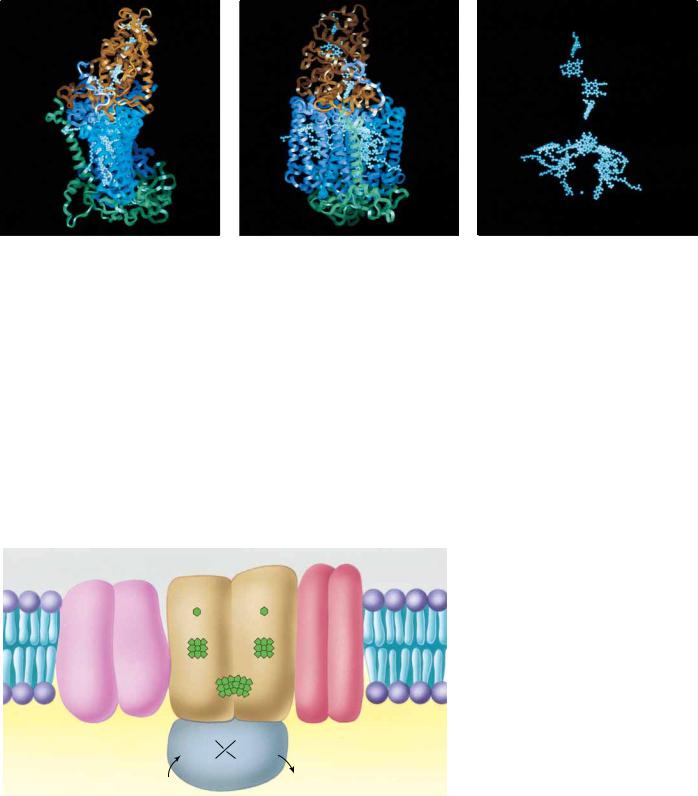
FIGURE 22.18 ● Model of the R. viridis reaction center. (a, b) Two views of the ribbon diagram of the reaction center. M and L subunits appear in purple and blue, respectively. Cytochrome subunit is brown; H subunit is green. These proteins provide a scaffold upon which the prosthetic groups of the reaction center are situated for effective photosynthetic electron transfer. Panel (c) shows the spatial relationship between the various prosthetic groups (4 hemes, P870, 2 BChl, 2 BPheo, 2 quinones, and the Fe atom) in the same view as in (b), but with protein chains deleted.
2 H2O 2 PQ 4 h n O2 2 PQH2. A cytochrome species, cytochrome b559, composed of two polypeptides (4.4 kD and 9.3 kD), is associated with PSII; its function is as yet unclear. Two chlorophyll-binding proteins (47 and 43 kD) harvest light and deliver exciton energy to P680. A Mn-containing extrinsic membrane protein, the OEC or oxygen-evolving complex (whose principal subunits are 33-, 23-, and 16-kD polypeptides) is located on the lumenal side of the thylakoid membrane.
|
|
|
|
|
Stroma |
|
|
QB |
Fe |
9.3kD |
4.5kD |
|
|
QA |
|
Chlorophyll |
Pheo |
Pheo |
|
|
|
Cytb559 |
|
antenna |
|
|
|
|
|
|
|
|
proteins |
|
P680 |
|
|
|
|
|
|
|
43kD |
47kD |
D1 |
|
D2 |
|
|
|
|
|
|
|
38kD |
D 39.4kD |
|
|
|
Mn |
Mn |
|
Thylakoid lumen |
|
|
Mn |
Mn |
|
|
|
2H2O |
OEC |
O2 |
|
FIGURE 22.19 ● The molecular architecture of PSII. The core of the PSII complex consists of the two polypeptides (D1 and D2) that bind P680, pheophytin (Pheo), and the quinones, Q A and Q B. Additional components of this complex include cytochrome b559, two additional intrinsic proteins (47 and 43 kD) that serve an accessory light-harvesting function, and an extrinsic protein complex that is essential to O2 evolution.
22.7 ● Light-Driven ATP Synthesis—Photophosphorylation |
727 |
Photosynthetic Energy Requirements for Hexose Synthesis
The fixation of carbon dioxide to form hexose, the dark reactions of photosynthesis, requires considerable energy. The overall stoichiometry of this process (Eq. 22.3) involves 12 NADPH and 18 ATP. To generate 12 equivalents of NADPH necessitates the consumption of 48 Einsteins of light, minimally 170 kJ each. However, if the preceding ratio of 113 ATP per NADPH were correct, insufficient ATP for CO2 fixation would be produced. Six additional Einsteins would provide the necessary two additional ATP. From 54 Einsteins, or 9180 kJ, one mole of hexose would be synthesized. The standard free energy change, G° , for hexose formation from carbon dioxide and water (the exact reverse of cellular respiration) is 2870 kJ/mol.
22.7 ● Light-Driven ATP Synthesis—Photophosphorylation
Light-driven ATP synthesis, termed photophosphorylation, is a fundamental part of the photosynthetic process. The conversion of light energy to chemical energy results in electron-transfer reactions leading to the generation of reducing power (NADPH). Coupled with these electron transfers, protons are driven across the thylakoid membranes from the stromal side to the lumenal side. These proton translocations occur in a manner analogous to the proton translocations accompanying mitochondrial electron transport that provide the driving force for oxidative phosphorylation (Chapter 21). Figure 22.12 indicates that proton translocations can occur at a number of sites. For example, protons may be translocated by reactions between H2O and PSII as a consequence of the photolysis of water. The oxidation–reduction events as electrons pass through the plastoquinone pool and the Q cycle are another source of proton translocations. The proton transfer accompanying NADP reduction also can be envisioned as protons being taken from the stromal side of the thylakoid vesicle. The current view is that two protons are translocated for each electron that flows from H2O to NADP . Because this electron transfer requires two photons, one falling at PSII and one at PSI, the overall yield is one proton per quantum of light.
The Mechanism of Photophosphorylation Is Chemiosmotic
The thylakoid membrane is asymmetrically organized, or “sided,” like the mitochondrial membrane. It also shares the property of being a barrier to the passive diffusion of H ions. Photosynthetic electron transport thus establishes an electrochemical gradient, or proton-motive force, across the thylakoid membrane with the interior, or lumen, side accumulating H ions relative to the stroma of the chloroplast. Like oxidative phosphorylation, the mechanism of photophosphorylation is chemiosmotic.
A proton-motive force of approximately 250 mV is needed to achieve ATP synthesis. This proton-motive force, p, is composed of a membrane potential, , and a pH gradient, pH (Chapter 21). The proton-motive force is defined as the free energy difference, G, divided by , Faraday’s constant:
In chloroplasts, the value of is typically 50 to 100 mV, and the pH gradient is equivalent to about 3 pH units, so that (2.3 RT/ ) pH 200 mV. This situation contrasts with the mitochondrial proton-motive force, where the membrane potential contributes relatively more to p than does the pH gradient.
728 Chapter 22 ● Photosynthesis
C R I T I C A L D E V E L O P M E N T S I N B I O C H E M I S T R Y
Experiments with Isolated Chloroplasts Provided the First Direct Evidence for the Chemiosmotic Hypothesis
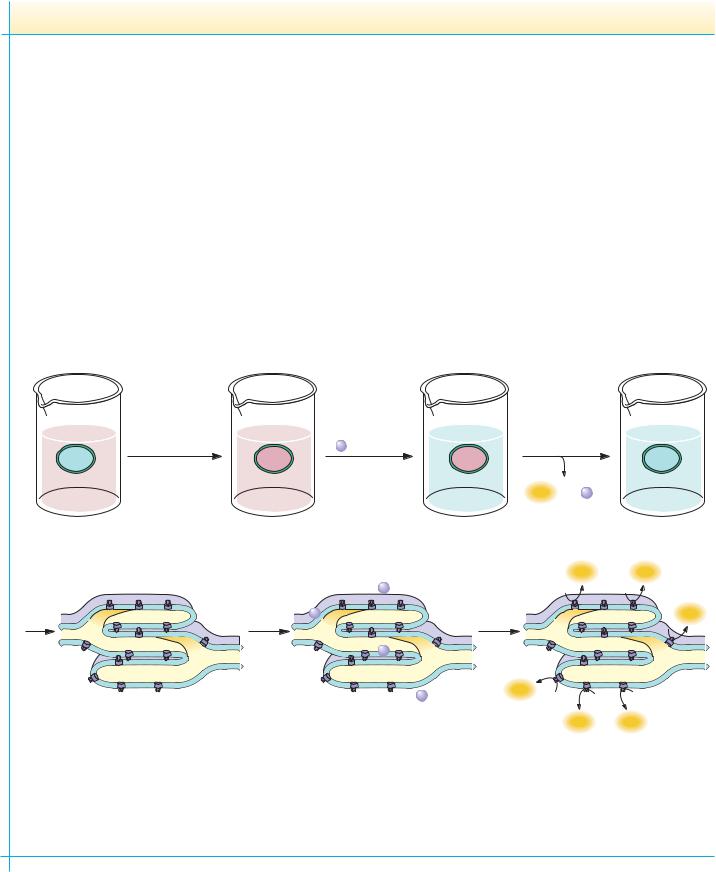
Experimental proof that the mechanism of photophosphorylation is chemiosmotic was provided in an elegant experiment by Andre Jagendorf and Ernest Uribe in 1966 (see figure). Jagendorf and Uribe reasoned that if photophosphorylation were indeed driven by an electrochemical gradient established by photosynthetic electron transfer reactions, they might artificially generate such a gradient by first incubating chloroplasts in an acid bath in the dark and then quickly raising the pH of the external medium. The resulting inequality in hydrogen ion electrochemical activity across the membrane should mimic the conditions normally found upon illumination of chloroplasts and should provide the energized condition necessary to drive ATP formation. To test this interpretation, Jagendorf and Uribe bathed isolated chloroplasts in a weakly acidic (pH 4) medium for 60 seconds, allowing the pH inside the chloroplasts to equilibrate with the external medium. The pH of the solution was then quickly raised to slightly
alkaline pH (pH 8), artificially creating a pH gradient across the thylakoid membranes. When ADP and Pi were added, ATP synthesis was observed as the pH gradient collapsed. This classic experiment was the first real proof of Mitchell’s chemiosmotic hypothesis and directed the scientific community to a greater acceptance of Mitchell’s interpretations. Mitchell’s chemiosmotic hypothesis for ATP synthesis now occupies the position of dogma as the weight of evidence has accumulated in its favor. Photophosphorylation then can be concisely summarized by noting that thylakoid vesicles accumulate H upon illumination and that the electrochemical gradient thus created represents an energized state that can be tapped to drive ATP synthesis. Collapse of the gradient—that is, equilibration of the ion concentration difference across the membrane—is the energy-transducing mechanism: the chemical potential of a concentration difference is transduced into synthesis of ATP.
|
|
Add ADP, |
|
60 sec |
|
32 P , and alkali |
15 sec |
pH 7 |
pH 4 |
pH 4 |
pH 8 |
pH 4 |
pH 4 |
pH 8 |
pH 8 |
|
|
|
ATP , γ -32 P |
|
|
|
labeled |
|
ADP |
|
|
ATP |
ATP |
|
P |
|
|
|
|
|
|
|
|
Acid |
Add |
|
ADP |
Followed |
ATP |
bath |
P |
|
by alkali |
ADP, Pi |
|
|
|
|
|
P |
|
|
|
|
ADP |
|
ADP |
ATP |
|
|
P |
|
|
|
|
|
|
|
|
|
|
|
ATP |
ATP |
The mechanism of photophosphorylation is chemiosmotic. In 1966, Jagendorf and Uribe experimentally demonstrated for the first time that establishment of an electrochemical gradient across the membrane of an energy-transducing organelle could lead to ATP synthesis. They equilibrated isolated chloroplasts for 60 seconds in a pH 4 bath, adjusted the pH to 8 in the presence of ADP and Pi, and allowed phosphorylation to proceed for 15 seconds. The entire experiment was carried out in the dark.
FIGURE 22.21
22.7 ● Light-Driven ATP Synthesis—Photophosphorylation |
729 |
CF1CF0 ATP Synthase Is the Chloroplast Equivalent of
the Mitochondrial F1F0 ATP Synthase
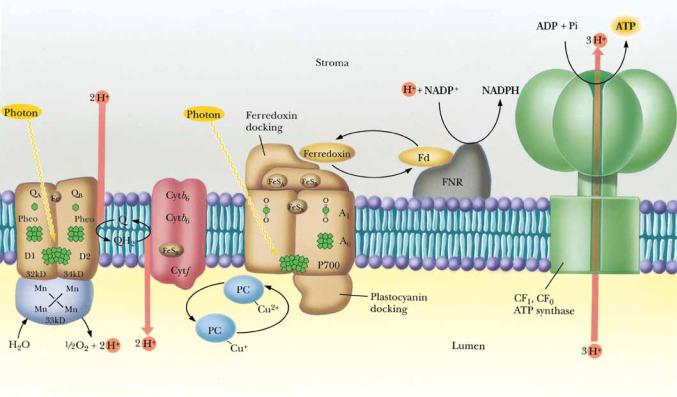
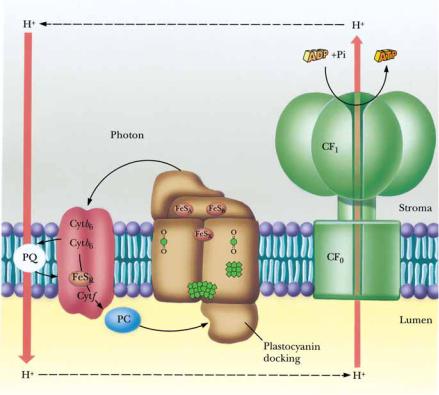
The transduction of the electrochemical gradient into the chemical energy represented by ATP is carried out by the chloroplast ATP synthase, which is highly analogous to the mitochondrial F1F0 ATP synthase. The chloroplast enzyme complex is called CF1CF0 ATP synthase, “C” symbolizing chloroplast. Like the mitochondrial complex, CF1CF0 ATP synthase is a heteromultimer of , , ,, , a, b, and c subunits (Chapter 21), consisting of a knoblike structure some 9 nm in diameter (CF1) attached to a stalked base (CF0) embedded in the thylakoid membrane. The mechanism of action of CF1CF0 ATP synthase in coupling ATP synthesis to the collapse of the pH gradient is similar to that of the mitochondrial ATP synthase described in Chapter 21. The mechanism of photophosphorylation is summarized schematically in Figure 22.21.
Cyclic and Noncyclic Photophosphorylation
Photosynthetic electron transport, which pumps H into the thylakoid lumen, can occur in two modes, both of which lead to the establishment of a transmembrane proton-motive force. Thus, both modes are coupled to ATP synthesis and are considered alternative mechanisms of photophosphorylation even though they are distinguished by differences in their electron transfer pathways. The two modes are cyclic and noncyclic photophosphorylation.
● The mechanism of photophosphorylation. Photosynthetic electron transport establishes a proton gradient that is tapped by the CF1CF0 ATP synthase to drive ATP synthesis. Critical to this mechanism is the fact that the membrane-bound components of light-induced electron transport and ATP synthesis are asymmetrical with respect to the thylakoid membrane so that vectorial discharge and uptake of H ensue, generating the proton-motive force.
730 Chapter 22 ● Photosynthesis
Noncyclic photophosphorylation has been the focus of our discussion and is represented by the scheme in Figure 22.21, where electrons activated by quanta at PSII and PSI flow from H2O to NADP , with concomitant establishment of the proton-motive force driving ATP synthesis. Note that in noncyclic photophosphorylation, O2 is evolved and NADP is reduced.
Cyclic Photophosphorylation
In cyclic photophosphorylation, the “electron hole” in P700 created by electron loss from P700 is filled not by an electron derived from H2O via PSII but by a cyclic pathway in which the photoexcited electron returns ultimately to P700 . This pathway is schematically represented in Figure 22.12 by the dashed line connecting FB and cytochrome b6. Thus, the function of cytochrome b6 (b563) is to couple the bound ferredoxin carriers of the PSI complex with the cytochrome b6/cytochrome f complex via the plastoquinone pool. This pathway diverts the activated e from NADP reduction back through plastocyanin to re-reduce P700 (Figure 22.22).
Proton translocations accompany these cyclic electron transfer events, so ATP synthesis can be achieved. In cyclic photophosphorylation, ATP is the sole product of energy conversion. No NADPH is generated, and, because PSII is not involved, no oxygen is evolved. The maximal rate of cyclic photophosphorylation is less than 5% of the rate of noncyclic photophosphorylation. Cyclic photophosphorylation depends only on PSI.
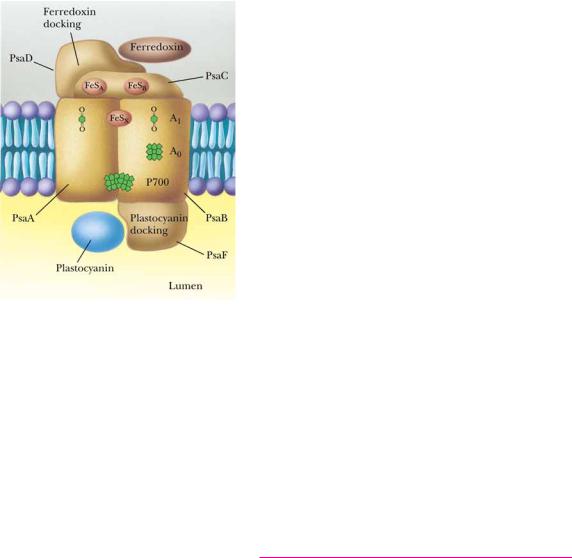
FIGURE 22.22 ● The pathway of cyclic photophosphorylation by PSI. (Adapted from Arnon,
D. I., 1984. Trends in Biochemical Sciences 9:258.)





 4 heme groups
4 heme groups
 H
H O C
O C  O
O 





 M
M 

 L
L