
Garrett R.H., Grisham C.M. - Biochemistry (1999)(2nd ed.)(en)
.pdf
102 Chapter 4 ● Amino Acids
encounter. All of these methods take advantage of the relative differences in the physical and chemical characteristics of amino acids, particularly ionization behavior and solubility characteristics. The methods important for amino acids include separations based on partition properties (the tendency to associate with one solvent or phase over another) and separations based on electrical charge. In all of the partition methods discussed here, the molecules of interest are allowed (or forced) to flow through a medium consisting of two phases—solid–liquid, liquid–liquid, or gas–liquid. In all of these methods, the molecules must show a preference for associating with one or the other phase. In this manner, the molecules partition, or distribute themselves, between the two phases in a manner based on their particular properties. The ratio of the concentrations of the amino acid (or other species) in the two phases is designated the partition coefficient.
In 1903, a separation technique based on repeated partitioning between phases was developed by Mikhail Tswett for the separation of plant pigments (carotenes and chlorophylls). Tswett, a Russian botanist, poured solutions of the pigments through columns of finely divided alumina and other solid media, allowing the pigments to partition between the liquid solvent and the solid support. Owing to the colorful nature of the pigments thus separated, Tswett called his technique chromatography. This term is now applied to a wide variety of separation methods, regardless of whether the products are colored or not. The success of all chromatography techniques depends on the repeated microscopic partitioning of a solute mixture between the available phases. The more frequently this partitioning can be made to occur within a given time span or over a given volume, the more efficient is the resulting separation. Chromatographic methods have advanced rapidly in recent years, due in part to the development of sophisticated new solid-phase materials. Methods important for amino acid separations include ion exchange chromatography, gas chromatography (GC), and high-performance liquid chromatography (HPLC).
Ion Exchange Chromatography
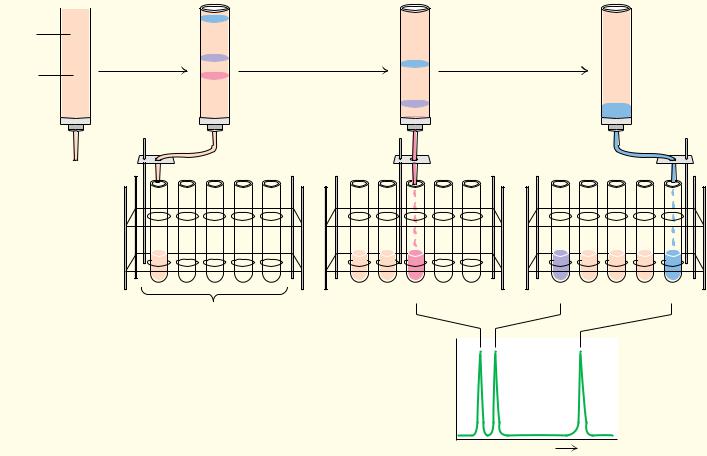
The separation of amino acids and other solutes is often achieved by means of ion exchange chromatography, in which the molecule of interest is exchanged for another ion onto and off of a charged solid support. In a typical procedure, solutes in a liquid phase, usually water, are passed through columns filled with a porous solid phase, usually a bed of synthetic resin particles, containing charged groups. Resins containing positive charges attract negatively charged solutes and are referred to as anion exchangers. Solid supports possessing negative charges attract positively charged species and are referred to as cation exchangers. Several typical cation and anion exchange resins with different types of charged groups are shown in Figure 4.18. The strength of the acidity or basicity of these groups and their number per unit volume of resin determine the type and strength of binding of an exchanger. Fully ionized acidic groups such as sulfonic acids result in an exchanger with a negative charge which binds cations very strongly. Weakly acidic or basic groups yield resins whose charge (and binding capacity) depends on the pH of the eluting solvent. The choice of the appropriate resin depends on the strength of binding desired. The bare charges on such solid phases must be counterbalanced by oppositely charged ions in solution (“counterions”). Washing a cation exchange resin, such as Dowex-50, which has strongly acidic phenyl-SO3 groups, with a NaCl solution results in the formation of the so-called sodium form of the resin (see Figure 4.19). When the mixture whose separation is desired is added to the column, the positively charged solute molecules displace the Na ions and bind to the



C |
hapter 5 |
. . . by small and simple things are great |
|
things brought to pass. |
|
|
ALMA 37.6, The Book of Mormon |
Proteins: Their Biological
Functions and Primary
Structure
OUTLINE
5.1 ● Proteins Are Linear Polymers
of Amino Acids
5.2 ● Architecture of Protein Molecules
5.3 ● The Many Biological Functions
of Proteins
5.4 ● Some Proteins Have Chemical Groups
Other Than Amino Acids
5.5 ● Reactions of Peptides and Proteins
5.6 ● Purification of Protein Mixtures
5.7 ● The Primary Structure of a Protein:
Determining the Amino Acid Sequence
5.8 ● Nature of Amino Acid Sequences
5.9 ● Synthesis of Polypeptides
in the Laboratory
Although helices are uncommon in manmade architecture, they are a common structural theme in biological macromole- cules—proteins, nucleic acids, and even polysaccharides.
(Loretto Chapel, Santa Fe, NM/ © Sarbo)
Proteins are a diverse and abundant class of biomolecules, constituting more than 50% of the dry weight of cells. This diversity and abundance reflect the central role of proteins in virtually all aspects of cell structure and function. An extraordinary diversity of cellular activity is possible only because of the versatility inherent in proteins, each of which is specifically tailored to its biological role. The pattern by which each is tailored resides within the genetic information of cells, encoded in a specific sequence of nucleotide bases in DNA.
107

108 Chapter 5 ● Proteins: Their Biological Functions and Primary Structure
Each such segment of encoded information defines a gene, and expression of the gene leads to synthesis of the specific protein encoded by it, endowing the cell with the functions unique to that particular protein. Proteins are the agents of biological function; they are also the expressions of genetic information.
5.1 ● Proteins Are Linear Polymers of Amino Acids
Chemically, proteins are unbranched polymers of amino acids linked head to tail, from carboxyl group to amino group, through formation of covalent peptide bonds, a type of amide linkage (Figure 5.1).
Peptide bond formation results in the release of H2O. The peptide “backbone” of a protein consists of the repeated sequence ONOC OCO, where the N represents the amide nitrogen, the C is the -carbon atom of an amino acid in the polymer chain, and the final C is the carbonyl carbon of the amino acid, which in turn is linked to the amide N of the next amino acid down the line. The geometry of the peptide backbone is shown in Figure 5.2. Note that the carbonyl oxygen and the amide hydrogen are trans to each other in this figure. This conformation is favored energetically because it results in less steric hindrance between nonbonded atoms in neighboring amino acids. Because the -carbon atom of the amino acid is a chiral center (in all amino acids except glycine), the polypeptide chain is inherently asymmetric. Only L-amino acids are found in proteins.
The Peptide Bond Has Partial Double Bond Character
The peptide linkage is usually portrayed by a single bond between the carbonyl carbon and the amide nitrogen (Figure 5.3a). Therefore, in principle, rotation may occur about any covalent bond in the polypeptide backbone because all three kinds of bonds (NOC , C OCo, and the CoON peptide bond) are single bonds. In this representation, the C and N atoms of the peptide grouping are both in planar sp2 hybridization and the C and O atoms are linked by a bond, leaving the nitrogen with a lone pair of electrons in a 2p orbital. However, another resonance form for the peptide bond is feasible in which the C and N atoms participate in a bond, leaving a lone e pair on the oxygen (Figure 5.3b). This structure prevents free rotation about the CoON peptide bond because it becomes a double bond. The real nature of the peptide bond lies somewhere between these extremes; that is, it has partial double bond character, as represented by the intermediate form shown in Figure 5.3c.
Peptide bond resonance has several important consequences. First, it restricts free rotation around the peptide bond and leaves the peptide backbone with only two degrees of freedom per amino acid group: rotation around
|
|
R1 |
O |
|
|
|
R2 |
O |
|
|
R1 |
|
O |
|
|
|
R2 |
O |
||||||||||
+ |
|
|
|
|
|
+ |
+ |
|
|
|
|
|
|
|
+ |
|
|
|
|
|
|
|
|
|
|
|||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
||||
H3N |
|
CH |
|
C |
O– |
H3N |
|
CH |
|
C |
O– |
|
|
H3N CH |
|
C |
|
N |
|
CH |
|
C |
O– |
|||||
|
|
|
|
|
|
|
|
|
|
|||||||||||||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
H2O |
|
|
|
|
|
H |
||||||||
Amino acid 1 |
|
|
Amino acid 2 |
|
|
|
|
|
Dipeptide |
|
||||||||||||||||||
● Peptide formation is the creation of an amide bond between the carboxyl group of one amino acid and the amino group of another amino acid. R1 and R2 represent the R groups of two different amino acids.



 containing
containing  several
several
