16-09-2014_09-00-17 / fer-restriction
.doc|
MoBio Contents |
DNA Recombination Technology |
|
Chapter 9 |
A | B | C | D | E | F | G | H | I | J | K |
|
|
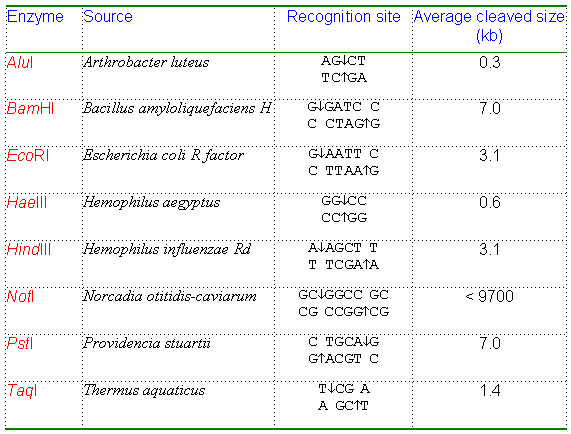
The procedure used in artificial DNA recombination is similar to the natural transpositional recombination. The major difference is that researchers can choose any appropriate enzymes to cut the DNA molecules. They are usually isolated from bacteria. The role of these enzymes in bacteria is to "restrict" the invasion of foreign DNA by cutting it into pieces. Hence, these enzymes are known as restriction enzymes. Table 9-A-1. Commonly used restriction enzymes.
Note: The "Recognition site" of a restriction enzyme is also called the restriction site. In this column, the first line is from 5' to 3' and the second line is from 3' to 5'. The arrow indicates the cleavage site. If the cleavage site is not at the center, the restriction enzyme will generate sticky ends (web link) which can base-pair with other DNA fragments cleaved by the same restriction enzyme. If the cleavage site is at the center, the restriction enzyme will generate blunt ends.
Illustrations from Genentech's Access Excellence The action of a restriction enzyme Inserting DNA into a plasmid Cloning into a plasmid
Other sites of Interest: REBASE - The restriction enzyme database. Web Cutter - a free web service for generating restriction maps (the map of restriction sties in a DNA sequence).
|