
Лекция4_2013_Комплемент
.pdf
( )
3b)nBbP
|
|
|
|
|
+ 6 |
+ 7 |
|
+ 8 |
|
+ 9 |
|||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
5 |
|
|
5b |
|
|
5b6 |
|
|
67 |
|
|
678 |
|||
|
|
|
|
|
|
|
|
|
|
||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|||
4b2 3b |
|
|
|
|
|
|
|
|
|
|
|
6789 |
|
||
C5a,
,
5b C6 C7 C8 C9-
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
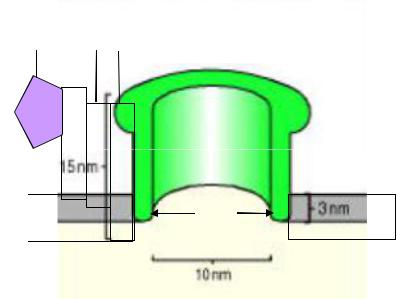
10-15 |
|
|
|
|
|
|
|
|
|
|
|||
|
|
|
|
|
|
|||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
-
I
S  C
C  O
O
II |
3b |
|
|
|
|
|
III |
3c |
|
|
|
|
|
|
|
|
|
|
|
|
|
||||
|
|
|
|
|
|
|
|
|
||||
|
|
|
|
|
|
|
|
|
||||
|
|
|
|
|
|
|
|
|
||||
|
|
|
|
|
|
|
|
|
|
|
||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
S C=O
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
3dg |
|
|||
3f
C3-
|
|
|
|
|
|
|
3b |
|
|
|
|
||
|
|
HS C=O |
3g |
|||
|
|
|
|
|||
|
|
O |
|
|||
|
|
|
|
|
||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
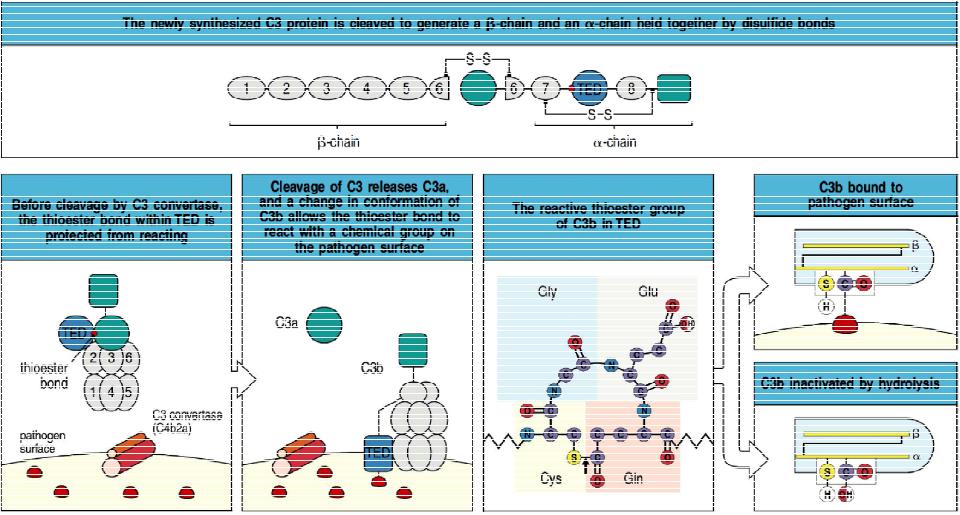
C3 convertase activates C3 for covalent bonding to microbial surfaces by cleaving it into C3a and C3b and exposing a highly reactive thioester bond in C3b.Top panel: C3 in blood plasma consists of an a-chain and a b-chain (formed by proteolytic processing from the native C3 polypeptide) held together by a disulfide bond. T he thioester-containing domain (fED) of the a-chain contains a potentially highly reactive thioester bond (red spot). Bottom left panels: cleavage by C3 convertase (the lectin pathway convertase C4b2a is shown here) and release of C3a from the amino terminus of the a-chain causes a conformational change in C3b that exposes the thioester bond. This can now react with hydroxyl or amino groups on molecules on microbial surfaces, covalently bonding C3b to the surface. Bottom right panels: schematic view of the thioester reaction. If a bond is not made with a microbial surface, the thioester is rapidly hydrolyzed (that is, cleaved by water), rendering C3b inactive.
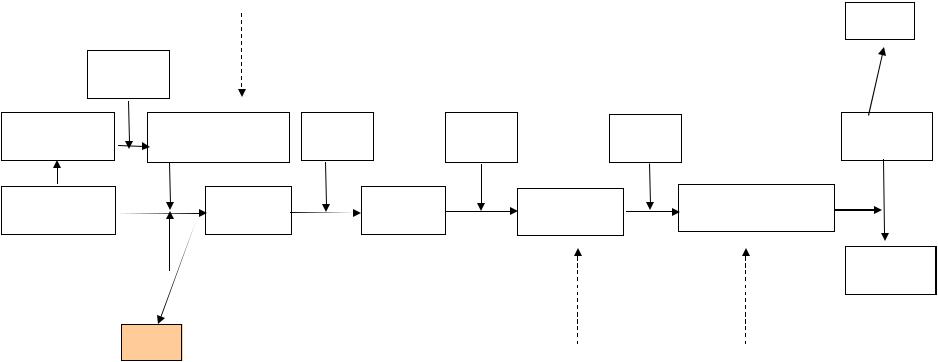
3-
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
,D |
|
|
|
|
|
|
|
|
||||||||||
|
|
|
|
|
|
|
|
|
|
|
|||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
3( 2 ) |
|
|
3( 2 )Bb |
|
|
|
|
.D |
|
|
|
|
|
5 |
|
||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
3 |
|
|
|
|
3b |
|
|
3bB |
|
3bBb |
|
3b)nBbP |
|
|
|
|
|||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
5b |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
3- 5-
|
,D |
|
|
|
|
|
|
|
|
|
|||||||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
3( 2 ) |
|
|
3( 2 )Bb |
|
|
|
|
D |
|
|
|
|
|
|
|
||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
3b |
|
|
3bB |
|
|
3bBb |
|
3b)nBbP |
|
|
|||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|||||||||||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
3 |
5 |
5b |
4b2a3b
|
|
|
|
|
|
MBL-MASP |
|
|
|
|
|
||
|
MBL |
|
|
|
|
|
|
|
|||||
|
|
|
|
|
|
|
|
|
|
|
|
||
|
|
|
MASP-1,2,3 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
4 |
|
|
4b |
|
|
|
4b2a |
||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
1q |
|
|
|
1qr |
2 |
|
||||||
|
|
|
|
|
|
s |
|
|
|
|
|
||
|
|
|
|
|
|
|
|
|
|
|
|||
|
|
|
|
|
|
|
|
|
5b6 |
|
67 |
|
678 |
|
|
||
|
|
|
6789 |
|
|
|||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
- 1rs |
|
-
-
3 5
|
|
|
|
|
|
|
|
|
|
|
|
- |
|
|
|
|
|
||||
|
|
|
|
|
|
|
|
|
||
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|||
|
|
|
|
|
|
|
||||
|
|
|
|
|
|
|
|
|
||
|
3 |
|
|
|
– |
|
|
|
||
|
|
|
|
|
|
|
|
|
|
|
4b 2a 4bC2aC3b
3bBb
|
|
. |
|
|
|
3b C5b, C3a C5a |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
- |
||
|
|||
|
|

1.C1- ( 1inh)
3-
2.C4- ( 4bp)
3.I ( 3b-INA)
5.3- CR1
6.MCP (Membrane cofactor protein).
7.DAF (Decay-activating factor - CD55)
8.S ( )
9.(C8-binding protein)
10.CD59 ( )
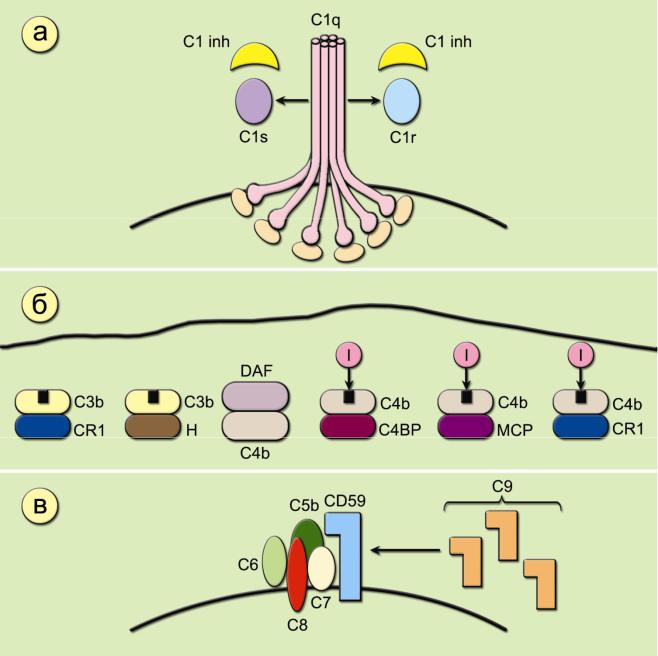
1.: ,
.
2.: C1->C4->C2->C3->C5->C6->C7->C8->C9.
3..
– 3 . , – ,
.
4..
.
. ( . )
« ».
(2 , )
).
