Embedded Robotics (Thomas Braunl, 2 ed, 2006)
.pdf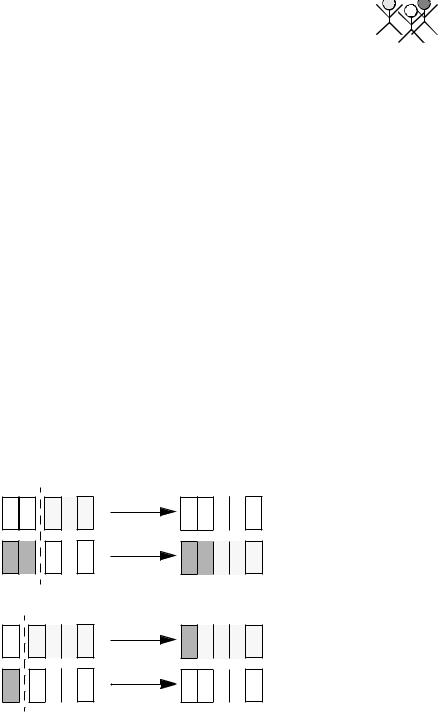


Implementation of Genetic Algorithms
x |
Bit String |
f(x) |
Ranking |
|
|
|
|
|
|
|
|
4 |
00100 |
–4 |
1 |
|
|
|
|
8 |
01000 |
–4 |
2 |
|
|
|
|
3 |
00011 |
–9 |
3 |
|
|
|
|
2 |
00010 |
–16 |
4 |
|
|
|
|
0 |
00000 |
–36 |
5 |
|
|
|
|
Table 20.3: Connection between the input and output indices
x = 6. This is because the x = 6 chromosome is now persistent through subsequent populations due to its optimal nature. When another chromosome is set to x = 6 through crossover, the chance of it being preserved through populations increases due to its increased presence in the population. This probability is proportional to the presence of the x = 6 chromosome in the population, and hence given enough iterations the whole population should converge. The elitism operator, combined with the fact that there is only one maximum, ensures that the population will never converge to another chromosome.
20.5 Implementation of Genetic Algorithms
We have implemented a genetic algorithm framework in object-oriented C++ for the robot projects described in the following chapters. The base system consists of abstract classes Gene, Chromosome, and Population. These classes may be extended with the functionality to handle different data types, including the advanced operators described earlier and for use in other applications as required. The implementation has been kept simple to meet the needs of the application it was developed for. More fully featured third-party genetic algorithm libraries are also freely available for use in complex applications, such as GA Lib [GALib 2006] and OpenBeagle [Beaulieu, Gagné 2006]. These allow us to begin designing a working genetic algorithm without having to implement any infrastructure. The basic relationship between program classes in these frameworks tends to be similar.
Using C++ and an object-oriented methodology maps well to the individual components of a genetic algorithm, allowing us to represent components by classes and operations by class methods. The concept of inheritance allows the base classes to be extended for specific applications without modification of the original code.
The basic unit of the system is a child of the base Gene class. Each instance of a Gene corresponds to a single parameter. The class itself is completely abstract: there is no default implementation, hence it is more accurately described as an interface. The Gene interface describes a set of basic opera-
301